Secretome analysis reveals effector candidates associated with broad host range necrotrophy in the fungal plant pathogen Sclerotinia sclerotiorum
- PMID: 24886033
- PMCID: PMC4039746
- DOI: 10.1186/1471-2164-15-336
Secretome analysis reveals effector candidates associated with broad host range necrotrophy in the fungal plant pathogen Sclerotinia sclerotiorum
Abstract
Background: The white mold fungus Sclerotinia sclerotiorum is a devastating necrotrophic plant pathogen with a remarkably broad host range. The interaction of necrotrophs with their hosts is more complex than initially thought, and still poorly understood.
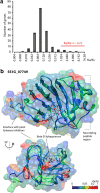
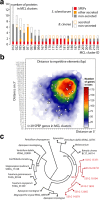
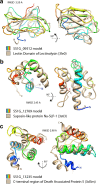
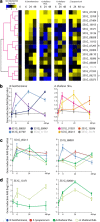
Results: We combined bioinformatics approaches to determine the repertoire of S. sclerotiorum effector candidates and conducted detailed sequence and expression analyses on selected candidates. We identified 486 S. sclerotiorum secreted protein genes expressed in planta, many of which have no predicted enzymatic activity and may be involved in the interaction between the fungus and its hosts. We focused on those showing (i) protein domains and motifs found in known fungal effectors, (ii) signatures of positive selection, (iii) recent gene duplication, or (iv) being S. sclerotiorum-specific. We identified 78 effector candidates based on these properties. We analyzed the expression pattern of 16 representative effector candidate genes on four host plants and revealed diverse expression patterns.
Conclusions: These results reveal diverse predicted functions and expression patterns in the repertoire of S. sclerotiorum effector candidates. They will facilitate the functional analysis of fungal pathogenicity determinants and should prove useful in the search for plant quantitative disease resistance components active against the white mold.
Figures



Similar articles
-
An Interspecies Comparative Analysis of the Predicted Secretomes of the Necrotrophic Plant Pathogens Sclerotinia sclerotiorum and Botrytis cinerea.PLoS One. 2015 Jun 24;10(6):e0130534. doi: 10.1371/journal.pone.0130534. eCollection 2015. PLoS One. 2015. PMID: 26107498 Free PMC article.
-
Draft genome sequencing and secretome profiling of Sclerotinia sclerotiorum revealed effector repertoire diversity and allied broad-host range necrotrophy.Sci Rep. 2022 Dec 17;12(1):21855. doi: 10.1038/s41598-022-22028-z. Sci Rep. 2022. PMID: 36528657 Free PMC article.
-
Genome-wide alternative splicing profiling in the fungal plant pathogen Sclerotinia sclerotiorum during the colonization of diverse host families.Mol Plant Pathol. 2021 Jan;22(1):31-47. doi: 10.1111/mpp.13006. Epub 2020 Oct 28. Mol Plant Pathol. 2021. PMID: 33111422 Free PMC article.
-
The evolutionary and molecular features of the broad-host-range plant pathogen Sclerotinia sclerotiorum.Mol Plant Pathol. 2022 Aug;23(8):1075-1090. doi: 10.1111/mpp.13221. Epub 2022 Apr 11. Mol Plant Pathol. 2022. PMID: 35411696 Free PMC article. Review.
-
Pathogenic attributes of Sclerotinia sclerotiorum: switching from a biotrophic to necrotrophic lifestyle.Plant Sci. 2015 Apr;233:53-60. doi: 10.1016/j.plantsci.2014.12.018. Epub 2014 Dec 31. Plant Sci. 2015. PMID: 25711813 Review.
Cited by
-
Elucidating the Role of Effectors in Plant-Fungal Interactions: Progress and Challenges.Front Microbiol. 2016 Apr 27;7:600. doi: 10.3389/fmicb.2016.00600. eCollection 2016. Front Microbiol. 2016. PMID: 27199930 Free PMC article. Review.
-
Population and genome-wide association studies of Sclerotinia sclerotiorum isolates collected from diverse host plants throughout the United States.Front Microbiol. 2023 Sep 27;14:1251003. doi: 10.3389/fmicb.2023.1251003. eCollection 2023. Front Microbiol. 2023. PMID: 37829452 Free PMC article.
-
Surface frustration re-patterning underlies the structural landscape and evolvability of fungal orphan candidate effectors.Nat Commun. 2023 Aug 28;14(1):5244. doi: 10.1038/s41467-023-40949-9. Nat Commun. 2023. PMID: 37640704 Free PMC article.
-
Label free proteomics and systematic analysis of secretome reveals effector candidates regulated by SGE1 and FTF1 in the plant pathogen Fusarium oxysporum f. sp. cubense tropical race 4.BMC Genomics. 2020 Apr 3;21(1):275. doi: 10.1186/s12864-020-6695-9. BMC Genomics. 2020. PMID: 32245409 Free PMC article.
-
Comparative secretome analysis of Striga and Cuscuta species identifies candidate virulence factors for two evolutionarily independent parasitic plant lineages.BMC Plant Biol. 2024 Apr 6;24(1):251. doi: 10.1186/s12870-024-04935-7. BMC Plant Biol. 2024. PMID: 38582844 Free PMC article.
References
-
- Peltier AJ, Bradley CA, Chilvers MI, Malvick DK, Mueller DS, Wise KA, Esker PD. Biology, yield loss and control of Sclerotinia stem rot of soybean. J Integrated Pest Manage. 2012;3(2):B1–B7. doi: 10.1603/IPM11033. - DOI
-
- Raffaele S, Kamoun S. Genome evolution in filamentous plant pathogens: why bigger can be better. Nat Rev Microbiol. 2012;10(6):417–430. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

