Laboratory evolution of adenylyl cyclase independent learning in Drosophila and missing heritability
- PMID: 24888634
- PMCID: PMC4108996
- DOI: 10.1111/gbb.12146
Laboratory evolution of adenylyl cyclase independent learning in Drosophila and missing heritability
Abstract
Gene interactions are acknowledged to be a likely source of missing heritability in large-scale genetic studies of complex neurological phenotypes. However, involvement of rare variants, de novo mutations, genetic lesions that are not easily detected with commonly used methods and epigenetic factors also are possible explanations. We used a laboratory evolution study to investigate the modulatory effects of background genetic variation on the phenotypic effect size of a null mutation with known impact on olfactory learning. To accomplish this, we first established a population that contained variation at just 23 loci and used selection to evolve suppression of the learning defect seen with null mutations in the rutabaga adenylyl cyclase. We thus biased the system to favor relatively simplified outcomes by choosing a Mendelian trait and by restricting the genetic variation segregating in the population. This experimental design also assures that the causal effects are among the known 23 segregating loci. We observe a robust response to selection that requires the presence of the 23 variants. Analyses of the underlying genotypes showed that interactions between more than two loci are likely to be involved in explaining the selection response, with implications for the missing heritability problem.
Keywords: Adenylyl cyclase; Drosophila; cAMP; cryptic variation; epistasis; laboratory evolution; learning; memory; missing heritability; selective breeding.
© 2014 John Wiley & Sons Ltd and International Behavioural and Neural Genetics Society.
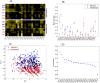
Figures


Similar articles
-
A neurofibromatosis-1-regulated pathway is required for learning in Drosophila.Nature. 2000 Feb 24;403(6772):895-8. doi: 10.1038/35002593. Nature. 2000. PMID: 10706287
-
The Drosophila learning and memory gene rutabaga encodes a Ca2+/Calmodulin-responsive adenylyl cyclase.Cell. 1992 Feb 7;68(3):479-89. doi: 10.1016/0092-8674(92)90185-f. Cell. 1992. PMID: 1739965
-
Spatiotemporal rescue of memory dysfunction in Drosophila.Science. 2003 Dec 5;302(5651):1765-8. doi: 10.1126/science.1089035. Science. 2003. PMID: 14657498
-
Drosophila bristles and the nature of quantitative genetic variation.Philos Trans R Soc Lond B Biol Sci. 2005 Jul 29;360(1459):1513-27. doi: 10.1098/rstb.2005.1672. Philos Trans R Soc Lond B Biol Sci. 2005. PMID: 16108138 Free PMC article. Review.
-
Physiology and biochemistry of Drosophila learning mutants.Physiol Rev. 1996 Apr;76(2):299-317. doi: 10.1152/physrev.1996.76.2.299. Physiol Rev. 1996. PMID: 8618959 Review.
References
-
- Alberini CM. Genes to remember. J Exp Biol. 1999;202:2887–91. - PubMed
-
- Barton NH, Keightley PD. Understanding quantitative genetic variation. Nat Rev Genet. 2002;3:11–21. - PubMed
-
- Daan S, Spoelstra K, Albrecht U, Schmutz I, Daan M, Daan B, Rienks F, Poletaeva I, Dell'omo G, Vyssotski A, Lipp HP. Lab mice in the field: unorthodox daily activity and effects of a dysfunctional circadian clock allele. J Biol Rhythms. 2011;26:118–29. - PubMed
-
- Davis RL. Olfactory memory formation in Drosophila: from molecular to systems neuroscience. Annu Rev Neurosci. 2005;28:275–302. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

