ATP-dependent chromatin remodeling complexes as novel targets for cancer therapy
- PMID: 24889532
- PMCID: PMC4839282
- DOI: 10.1016/B978-0-12-800249-0.00005-6
ATP-dependent chromatin remodeling complexes as novel targets for cancer therapy
Abstract
The progression to advanced stage cancer requires changes in many characteristics of a cell. These changes are usually initiated through spontaneous mutation. As a result of these mutations, gene expression is almost invariably altered allowing the cell to acquire tumor-promoting characteristics. These abnormal gene expression patterns are in part enabled by the posttranslational modification and remodeling of nucleosomes in chromatin. These chromatin modifications are established by a functionally diverse family of enzymes including histone and DNA-modifying complexes, histone deposition pathways, and chromatin remodeling complexes. Because the modifications these enzymes deposit are essential for maintaining tumor-promoting gene expression, they have recently attracted much interest as novel therapeutic targets. One class of enzyme that has not generated much interest is the chromatin remodeling complexes. In this review, we will present evidence from the literature that these enzymes have both causal and enabling roles in the transition to advanced stage cancers; as such, they should be seriously considered as high-value therapeutic targets. Previously published strategies for discovering small molecule regulators to these complexes are described. We close with thoughts on future research, the field should perform to further develop this potentially novel class of therapeutic target.
Keywords: Cancer; Chromatin; Chromatin remodeling; Epigenetics; Nucleosome.
© 2014 Elsevier Inc. All rights reserved.
Figures

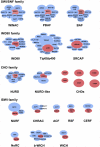

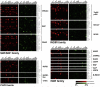
Similar articles
-
Mechanisms for nucleosome movement by ATP-dependent chromatin remodeling complexes.Results Probl Cell Differ. 2006;41:127-48. doi: 10.1007/400_005. Results Probl Cell Differ. 2006. PMID: 16909894 Review.
-
ATP-dependent nucleosome remodeling complexes: enzymes tailored to deal with chromatin.J Cell Biochem. 2004 Apr 15;91(6):1087-98. doi: 10.1002/jcb.20005. J Cell Biochem. 2004. PMID: 15048866 Review.
-
Chromatin remodeling by imitation switch (ISWI) class ATP-dependent remodelers is stimulated by histone variant H2A.Z.J Biol Chem. 2010 Feb 12;285(7):4645-51. doi: 10.1074/jbc.M109.072348. Epub 2009 Nov 25. J Biol Chem. 2010. PMID: 19940112 Free PMC article.
-
A novel mechanism of antagonism between ATP-dependent chromatin remodeling complexes regulates RNR3 expression.Mol Cell Biol. 2009 Jun;29(12):3255-65. doi: 10.1128/MCB.01741-08. Epub 2009 Apr 6. Mol Cell Biol. 2009. PMID: 19349301 Free PMC article.
-
The ATP-dependent chromatin remodeling enzyme Fun30 represses transcription by sliding promoter-proximal nucleosomes.J Biol Chem. 2013 Aug 9;288(32):23182-93. doi: 10.1074/jbc.M113.471979. Epub 2013 Jun 18. J Biol Chem. 2013. PMID: 23779104 Free PMC article.
Cited by
-
New inhibitors for the BPTF bromodomain enabled by structural biology and biophysical assay development.Org Biomol Chem. 2020 Jul 15;18(27):5174-5182. doi: 10.1039/d0ob00506a. Org Biomol Chem. 2020. PMID: 32588860 Free PMC article.
-
BPTF Is Essential for T Cell Homeostasis and Function.J Immunol. 2016 Dec 1;197(11):4325-4333. doi: 10.4049/jimmunol.1600642. Epub 2016 Oct 31. J Immunol. 2016. PMID: 27799308 Free PMC article.
-
SHU00238 Promotes Colorectal Cancer Cell Apoptosis Through miR-4701-3p and miR-4793-3p.Front Genet. 2020 Jan 10;10:1320. doi: 10.3389/fgene.2019.01320. eCollection 2019. Front Genet. 2020. PMID: 31998373 Free PMC article.
-
Epigenetic Targeting of Glioblastoma.Front Oncol. 2018 Oct 16;8:448. doi: 10.3389/fonc.2018.00448. eCollection 2018. Front Oncol. 2018. PMID: 30386738 Free PMC article. Review.
-
Autophagy-Dependent Sensitization of Triple-Negative Breast Cancer Models to Topoisomerase II Poisons by Inhibition of the Nucleosome Remodeling Factor.Mol Cancer Res. 2021 Aug;19(8):1338-1349. doi: 10.1158/1541-7786.MCR-20-0743. Epub 2021 Apr 2. Mol Cancer Res. 2021. PMID: 33811160 Free PMC article.
References
-
- Aalfs JD, Kingston RE. What does ‘chromatin remodeling’ mean? Trends in Biochemical Sciences. 2000;25(11):548–555. - PubMed
-
- Adams ME, Hurd EA, Beyer LA, Swiderski DL, Raphael Y, Martin DM. Defects in vestibular sensory epithelia and innervation in mice with loss of Chd7 function: Implications for human CHARGE syndrome. The Journal of Comparative Neurology. 2007;504(5):519–532. - PubMed
-
- Allis CD, Jenuwein T, Reinberg D. Epigenetics. Cold Spring Harbor Laboratory Press; Cold Spring Harbor, N.Y.: 2007.
-
- Arrowsmith CH, Bountra C, Fish PV, Lee K, Schapira M. Epigenetic protein families: A new frontier for drug discovery. Nature Reviews Drug Discovery. 2012;11(5):384–400. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

