Exploring the substrate range of wild-type aminoacyl-tRNA synthetases
- PMID: 24890918
- PMCID: PMC4133344
- DOI: 10.1002/cbic.201402083
Exploring the substrate range of wild-type aminoacyl-tRNA synthetases
Abstract
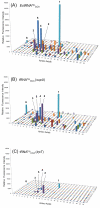
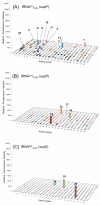
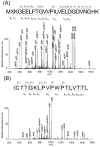
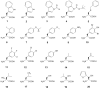
We tested the substrate range of four wild-type E. coli aminoacyl-tRNA synthetases (AARSs) with a library of nonstandard amino acids (nsAAs). Although these AARSs could discriminate efficiently against the other canonical amino acids, they were able to use many nsAAs as substrates. Our results also show that E. coli tryptophanyl-tRNA synthetase (TrpRS) and tyrosyl-tRNA synthetase have overlapping substrate ranges. In addition, we found that the nature of the anticodon sequence of tRNA(Trp) altered the nsAA substrate range of TrpRS; this implies that the sequence of the anticodon affects the TrpRS amino acid binding pocket. These results highlight again that inherent AARS polyspecificity will be a major challenge in the aim of incorporating multiple different amino acids site-specifically into proteins.
Keywords: aminoacyl-tRNA synthetases; natural nonsense tRNA suppressors; nonstandard amino acids; substrate specificity; synthetic biology; tRNA.
© 2014 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim.
Figures
References
-
- Richmond MH. Bacteriol Rev. 1962;26:398–420. - PMC - PubMed
- Xu ZJ, Love ML, Ma LY, Blum M, Bronskill PM, Bernstein J, Grey AA, Hofmann T, Camerman N, Wong JT. J Biol Chem. 1989;264:4304–4311. - PubMed
- Li L, Boniecki MT, Jaffe JD, Imai BS, Yau PM, Luthey-Schulten ZA, Martinis SA. Proc Natl Acad Sci USA. 2011;108:9378–9383. - PMC - PubMed
-
- Kavran JM, Gundllapalli S, O’Donoghue P, Englert M, Söll D, Steitz TA. Proc Natl Acad Sci U S A. 2007;104:11268–11273. - PMC - PubMed
- Ko JH, Wang YS, Nakamura A, Guo LT, Söll D, Umehara T. FEBS Lett. 2013;587:3243–3248. - PMC - PubMed
- Nguyen DP, Lusic H, Neumann H, Kapadnis PB, Deiters A, Chin JW. J Am Chem Soc. 2009;131:8720–8721. - PubMed
- Takimoto JK, Dellas N, Noel JP, Wang L. ACS Chem Biol. 2011;6:733–743. - PMC - PubMed
- Wang YS, Fang X, Chen HY, Wu B, Wang ZU, Hilty C, Liu WR. ACS Chem Biol. 2013;8:405–415. - PMC - PubMed
- Wang YS, Fang X, Wallace AL, Wu B, Liu WR. J Am Chem Soc. 2012;134:2950–2953. - PMC - PubMed
- Wang YS, Russell WK, Wang Z, Wan W, Dodd LE, Pai PJ, Russell DH, Liu WR. Mol Biosyst. 2011;7:714–717. - PubMed
- Yanagisawa T, Ishii R, Fukunaga R, Kobayashi T, Sakamoto K, Yokoyama S. Chem Biol. 2008;15:1187–1197. - PubMed
- Young DD, Young TS, Jahnz M, Ahmad I, Spraggon G, Schultz PG. Biochemistry. 2011;50:1894–1900. - PMC - PubMed
- Wang ZU, Wang YS, Pai PJ, Russell WK, Russell DH, Liu WR. Biochemistry. 2012;51:5232–5234. - PMC - PubMed
- Wang YS, Wu B, Wang Z, Huang Y, Wan W, Russell WK, Pai PJ, Moe YN, Russell DH, Liu WR. Mol Biosyst. 2010;6:1557–1560. - PubMed
- Fekner T, Li X, Lee MM, Chan MK. Angew Chem Int Ed Engl. 2009;48:1633–1635. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

