A chemical potentiator of copper-accumulation used to investigate the iron-regulons of Saccharomyces cerevisiae
- PMID: 24895027
- PMCID: PMC4149784
- DOI: 10.1111/mmi.12661
A chemical potentiator of copper-accumulation used to investigate the iron-regulons of Saccharomyces cerevisiae
Abstract
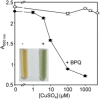
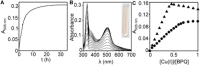
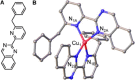
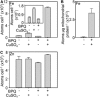
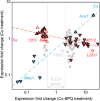
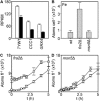
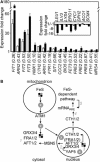
The extreme resistance of Saccharomyces cerevisiae to copper is overcome by 2-(6-benzyl-2-pyridyl)quinazoline (BPQ), providing a chemical-biology tool which has been exploited in two lines of discovery. First, BPQ is shown to form a red (BPQ)2 Cu(I) complex and promote Ctr1-independent copper-accumulation in whole cells and in mitochondria isolated from treated cells. Multiple phenotypes, including loss of aconitase activity, are consistent with copper-BPQ mediated damage to mitochondrial iron-sulphur clusters. Thus, a biochemical basis of copper-toxicity in S. cerevisiae is analogous to other organisms. Second, iron regulons controlled by Aft1/2, Cth2 and Yap5 that respond to mitochondrial iron-sulphur cluster status are modulated by copper-BPQ causing iron hyper-accumulation via upregulated iron-import. Comparison of copper-BPQ treated, untreated and copper-only treated wild-type and fra2Δ by RNA-seq has uncovered a new candidate Aft1 target-gene (LSO1) and paralogous non-target (LSO2), plus nine putative Cth2 target-transcripts. Two lines of evidence confirm that Fra2 dominates basal repression of the Aft1/2 regulons in iron-replete cultures. Fra2-independent control of these regulons is also observed but CTH2 itself appears to be atypically Fra2-dependent. However, control of Cth2-target transcripts which is independent of CTH2 transcript abundance or of Fra2, is also quantified. Use of copper-BPQ supports a substantial contribution of metabolite repression to iron-regulation.
© 2014 The Authors. Molecular Microbiology published by John Wiley & Sons Ltd.
Figures




Similar articles
-
The late-annotated small ORF LSO1 is a target gene of the iron regulon of Saccharomyces cerevisiae.Microbiologyopen. 2015 Dec;4(6):941-51. doi: 10.1002/mbo3.303. Epub 2015 Oct 8. Microbiologyopen. 2015. PMID: 26450372 Free PMC article.
-
Biophysical investigation of the iron in Aft1-1(up) and Gal-YAH1 Saccharomyces cerevisiae.Biochemistry. 2011 Apr 5;50(13):2660-71. doi: 10.1021/bi102015s. Epub 2011 Feb 28. Biochemistry. 2011. PMID: 21361388 Free PMC article.
-
Activation of the iron regulon by the yeast Aft1/Aft2 transcription factors depends on mitochondrial but not cytosolic iron-sulfur protein biogenesis.J Biol Chem. 2005 Mar 18;280(11):10135-40. doi: 10.1074/jbc.M413731200. Epub 2005 Jan 13. J Biol Chem. 2005. PMID: 15649888
-
Mechanisms of iron sensing and regulation in the yeast Saccharomyces cerevisiae.World J Microbiol Biotechnol. 2017 Apr;33(4):75. doi: 10.1007/s11274-017-2215-8. Epub 2017 Mar 17. World J Microbiol Biotechnol. 2017. PMID: 28315258 Review.
-
Iron sensing and regulation in Saccharomyces cerevisiae: Ironing out the mechanistic details.Curr Opin Microbiol. 2013 Dec;16(6):662-8. doi: 10.1016/j.mib.2013.07.020. Epub 2013 Aug 17. Curr Opin Microbiol. 2013. PMID: 23962819 Free PMC article. Review.
Cited by
-
Getting out what you put in: Copper in mitochondria and its impacts on human disease.Biochim Biophys Acta Mol Cell Res. 2021 Jan;1868(1):118867. doi: 10.1016/j.bbamcr.2020.118867. Epub 2020 Oct 2. Biochim Biophys Acta Mol Cell Res. 2021. PMID: 32979421 Free PMC article. Review.
-
Increasing the copper sensitivity of microorganisms by restricting iron supply, a strategy for bio-management practices.Microb Biotechnol. 2020 Sep;13(5):1530-1545. doi: 10.1111/1751-7915.13590. Epub 2020 Jun 19. Microb Biotechnol. 2020. PMID: 32558275 Free PMC article.
-
Copper at the Fungal Pathogen-Host Axis.J Biol Chem. 2015 Jul 31;290(31):18945-53. doi: 10.1074/jbc.R115.649129. Epub 2015 Jun 8. J Biol Chem. 2015. PMID: 26055724 Free PMC article. Review.
-
Reconstitution of a thermophilic Cu+ importer in vitro reveals intrinsic high-affinity slow transport driving accumulation of an essential metal ion.J Biol Chem. 2018 Oct 5;293(40):15497-15512. doi: 10.1074/jbc.RA118.004802. Epub 2018 Aug 21. J Biol Chem. 2018. PMID: 30131336 Free PMC article.
-
The late-annotated small ORF LSO1 is a target gene of the iron regulon of Saccharomyces cerevisiae.Microbiologyopen. 2015 Dec;4(6):941-51. doi: 10.1002/mbo3.303. Epub 2015 Oct 8. Microbiologyopen. 2015. PMID: 26450372 Free PMC article.
References
-
- Askwith C, Eide D, Van Ho A, Bernard PS, Li L, Davis-Kaplan S, et al. The FET3 gene of S. cerevisiae encodes a multicopper oxidase required for ferrous iron uptake. Cell. 1994;76:403–410. - PubMed
-
- Blaiseau PL, Lesuisse E, Camadro JM. Aft2p, a novel iron-regulated transcription activator that modulates, with Aft1p, intracellular iron use and resistance to oxidative stress in yeast. J Biol Chem. 2001;276:34221–34226. - PubMed
-
- Chen OS, Crisp RJ, Valachovic M, Bard M, Winge DR, Kaplan J. Transcription of the yeast iron regulon does not respond directly to iron but rather to iron-sulfur cluster biosynthesis. J Biol Chem. 2004;279:29513–29518. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

