Unraveling patterns of site-to-site synonymous rates variation and associated gene properties of protein domains and families
- PMID: 24896293
- PMCID: PMC4045579
- DOI: 10.1371/journal.pone.0095034
Unraveling patterns of site-to-site synonymous rates variation and associated gene properties of protein domains and families
Erratum in
- PLoS One. 2014;9(7):e102721
Abstract
In protein-coding genes, synonymous mutations are often thought not to affect fitness and therefore are not subject to natural selection. Yet increasingly, cases of non-neutral evolution at certain synonymous sites were reported over the last decade. To evaluate the extent and the nature of site-specific selection on synonymous codons, we computed the site-to-site synonymous rate variation (SRV) and identified gene properties that make SRV more likely in a large database of protein-coding gene families and protein domains. To our knowledge, this is the first study that explores the determinants and patterns of the SRV in real data. We show that the SRV is widespread in the evolution of protein-coding sequences, putting in doubt the validity of the synonymous rate as a standard neutral proxy. While protein domains rarely undergo adaptive evolution, the SRV appears to play important role in optimizing the domain function at the level of DNA. In contrast, protein families are more likely to evolve by positive selection, but are less likely to exhibit SRV. Stronger SRV was detected in genes with stronger codon bias and tRNA reusage, those coding for proteins with larger number of interactions or forming larger number of structures, located in intracellular components and those involved in typically conserved complex processes and functions. Genes with extreme SRV show higher expression levels in nearly all tissues. This indicates that codon bias in a gene, which often correlates with gene expression, may often be a site-specific phenomenon regulating the speed of translation along the sequence, consistent with the co-translational folding hypothesis. Strikingly, genes with SRV were strongly overrepresented for metabolic pathways and those associated with several genetic diseases, particularly cancers and diabetes.
Conflict of interest statement
Figures

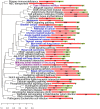

References
-
- Anfinsen CB (1973) Principles that govern the folding of protein chains. Science 181: 223–230. - PubMed
-
- Clarke B (1970) Darwinian evolution of proteins. Science 168: 1009–1011. - PubMed
-
- Ikemura T (1981) Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes. J Mol Biol 146: 1–21. - PubMed
-
- Akashi H, Eyre-Walker A (1998) Translational selection and molecular evolution. Curr Opin Genet Dev 8: 688–693. - PubMed
-
- Duret L (2002) Evolution of synonymous codon usage in metazoans. Curr Opin Genet Dev 12: 640–649. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

