Inhibitors of the Cdc34 acidic loop: A computational investigation integrating molecular dynamics, virtual screening and docking approaches
- PMID: 24918063
- PMCID: PMC4050183
- DOI: 10.1016/j.fob.2014.04.011
Inhibitors of the Cdc34 acidic loop: A computational investigation integrating molecular dynamics, virtual screening and docking approaches
Abstract
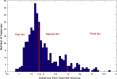
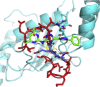
Among the different classes of enzymes involved in the ubiquitin pathway, E2 ubiquitin-conjugating enzymes occupy a central role in the ubiquitination cascade. Cdc34-like E2 enzymes are characterized by a 12-14 residue insertion in the proximity of the catalytic site, known as the acidic loop. Cdc34 ubiquitin-charging activity is regulated by CK2-dependent phosphorylation and the regulatory mechanism involves the acidic loop. Indeed, the phosphorylation stabilizes the loop in an open conformation that is competent for ubiquitin charging. Cdc34 is associated with a variety of diseases, such as hepatocellular carcinomas and prostatic adenocarcinomas. In light of its role, the discovery of potential inhibitory compounds would provide the mean to effectively modulate its activity. Here, we carried out a computational study based on molecular dynamics, virtual screening and docking to identify potential inhibitory compounds of Cdc34, modulating the acidic loop conformation. The molecules identified in this study have been designed to act as molecular hinges that can bind the acidic loop in its closed conformation, thus inhibiting the Cdc34-mediated ubiquitination cascade at the ubiquitin-charging step. In particular, we proposed a pharmacophore model featuring two amino groups in the central part of the model and two lateral aromatic chains, which respectively establish electrostatic interactions with the acidic loop (Asp 108 and Glu 109) and a hydrogen bond with Ser 139, which is one of the key residues for Cdc34 activity.
Keywords: Cdc34; Docking; E2 conjugating enzyme; MD, molecular dynamics; Sc, Saccharomyces cerevisiae; UBC, ubiquitin-binding domain; Ub, ubiquitin; Ubiquitin; Ubiquitination; Virtual screening.
Figures



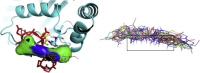





Similar articles
-
An acidic loop and cognate phosphorylation sites define a molecular switch that modulates ubiquitin charging activity in Cdc34-like enzymes.PLoS Comput Biol. 2011 May;7(5):e1002056. doi: 10.1371/journal.pcbi.1002056. Epub 2011 May 26. PLoS Comput Biol. 2011. PMID: 21637798 Free PMC article.
-
Multimodal mechanism of action for the Cdc34 acidic loop: a case study for why ubiquitin-conjugating enzymes have loops and tails.J Biol Chem. 2013 Nov 29;288(48):34882-96. doi: 10.1074/jbc.M113.509190. Epub 2013 Oct 15. J Biol Chem. 2013. PMID: 24129577 Free PMC article.
-
Molecular and structural insight into lysine selection on substrate and ubiquitin lysine 48 by the ubiquitin-conjugating enzyme Cdc34.Cell Cycle. 2013 Jun 1;12(11):1732-44. doi: 10.4161/cc.24818. Epub 2013 May 8. Cell Cycle. 2013. PMID: 23656784 Free PMC article.
-
Ubiquitin-proteasome system as a factor that determine the sensitivity to methylmercury.Yakugaku Zasshi. 2007 Mar;127(3):463-8. doi: 10.1248/yakushi.127.463. Yakugaku Zasshi. 2007. PMID: 17329932 Review.
-
The family of ubiquitin-conjugating enzymes (E2s): deciding between life and death of proteins.FASEB J. 2010 Apr;24(4):981-93. doi: 10.1096/fj.09-136259. Epub 2009 Nov 25. FASEB J. 2010. PMID: 19940261 Review.
Cited by
-
Structural Analysis, Multi-Conformation Virtual Screening and Molecular Simulation to Identify Potential Inhibitors Targeting pS273R Proteases of African Swine Fever Virus.Molecules. 2023 Jan 6;28(2):570. doi: 10.3390/molecules28020570. Molecules. 2023. PMID: 36677630 Free PMC article.
-
The Catalytically Inactive Mutation of the Ubiquitin-Conjugating Enzyme CDC34 Affects its Stability and Cell Proliferation.Protein J. 2018 Apr;37(2):132-143. doi: 10.1007/s10930-018-9766-x. Protein J. 2018. PMID: 29564676
-
Mechanism of Lysine 48 Selectivity during Polyubiquitin Chain Formation by the Ube2R1/2 Ubiquitin-Conjugating Enzyme.Mol Cell Biol. 2016 May 16;36(11):1720-32. doi: 10.1128/MCB.00097-16. Print 2016 Jun 1. Mol Cell Biol. 2016. PMID: 27044868 Free PMC article.
-
Ubiquitination Regulators Discovered by Virtual Screening for the Treatment of Cancer.Front Cell Dev Biol. 2021 May 12;9:665646. doi: 10.3389/fcell.2021.665646. eCollection 2021. Front Cell Dev Biol. 2021. PMID: 34055799 Free PMC article. Review.
-
E2 superfamily of ubiquitin-conjugating enzymes: constitutively active or activated through phosphorylation in the catalytic cleft.Sci Rep. 2015 Oct 14;5:14849. doi: 10.1038/srep14849. Sci Rep. 2015. PMID: 26463729 Free PMC article.
References
-
- Spasser L., Brik A. Chemistry and biology of the ubiquitin signal. Angew. Chem. Int. Ed. Engl. 2012;51:6840–6862. - PubMed
-
- Nagy V., Dikic I. Ubiquitin ligase complexes: from substrate selectivity to conjugational specificity. Biol. Chem. 2010;391:163–169. - PubMed
-
- Harper J.W., Schulman B.A. Structural complexity in ubiquitin recognition. Cell. 2006;124(2006):1133–1136. - PubMed
-
- Komander D., Rape M. The ubiquitin code. Annu. Rev. Biochem. 2012;81:203–229. - PubMed
-
- Komander D. The emerging complexity of protein ubiquitination. Biochem. Soc. Trans. 2009;37:937–953. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

