Production and characterization of high-titer serum-free cell culture grown hepatitis C virus particles of genotype 1-6
- PMID: 24928051
- PMCID: PMC4415741
- DOI: 10.1016/j.virol.2014.03.021
Production and characterization of high-titer serum-free cell culture grown hepatitis C virus particles of genotype 1-6
Abstract
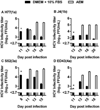
Recently, cell culture systems producing hepatitis C virus particles (HCVcc) were developed. Establishment of serum-free culture conditions is expected to facilitate development of a whole-virus inactivated HCV vaccine. We describe generation of genotype 1-6 serum-free HCVcc (sf-HCVcc) from Huh7.5 hepatoma cells cultured in adenovirus expression medium. Compared to HCVcc, sf-HCVcc showed 0.6-2.1 log10 higher infectivity titers (4.7-6.2 log10 Focus Forming Units/mL), possibly due to increased release and specific infectivity of sf-HCVcc. In contrast to HCVcc, sf-HCVcc had a homogeneous single-peak density profile. Entry of sf-HCVcc depended on HCV co-receptors CD81, LDLr, and SR-BI, and clathrin-mediated endocytosis. HCVcc and sf-HCVcc were neutralized similarly by chronic-phase patient sera and by human monoclonal antibodies targeting conformational epitopes. Thus, we developed serum-free culture systems producing high-titer single-density sf-HCVcc, showing similar biological properties as HCVcc. This methodology has the potential to advance HCV vaccine development and to facilitate biophysical studies of HCV.
Keywords: Adenovirus expression medium; Biophysical characterization; Cell culture system; Genotypes; Hepatitis C virus; High-titer; Neutralization; Receptor blocking; Serum-free; Vaccine development.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures











Similar articles
-
Adaptive Mutations Enhance Assembly and Cell-to-Cell Transmission of a High-Titer Hepatitis C Virus Genotype 5a Core-NS2 JFH1-Based Recombinant.J Virol. 2015 Aug;89(15):7758-75. doi: 10.1128/JVI.00039-15. Epub 2015 May 20. J Virol. 2015. PMID: 25995244 Free PMC article.
-
A serum protein factor mediates maturation and apoB-association of HCV particles in the extracellular milieu.J Hepatol. 2019 Apr;70(4):626-638. doi: 10.1016/j.jhep.2018.11.033. Epub 2018 Dec 14. J Hepatol. 2019. PMID: 30553840
-
Functional and Biochemical Characterization of Hepatitis C Virus (HCV) Particles Produced in a Humanized Liver Mouse Model.J Biol Chem. 2015 Sep 18;290(38):23173-87. doi: 10.1074/jbc.M115.662999. Epub 2015 Jul 29. J Biol Chem. 2015. PMID: 26224633 Free PMC article.
-
Similarities and Differences Between HCV Pseudoparticle (HCVpp) and Cell Culture HCV (HCVcc) in the Study of HCV.Methods Mol Biol. 2019;1911:33-45. doi: 10.1007/978-1-4939-8976-8_2. Methods Mol Biol. 2019. PMID: 30593616 Review.
-
Cutting the gordian knot-development and biological relevance of hepatitis C virus cell culture systems.Adv Virus Res. 2008;71:51-133. doi: 10.1016/S0065-3527(08)00002-X. Adv Virus Res. 2008. PMID: 18585527 Review.
Cited by
-
Adaptive Mutations Enhance Assembly and Cell-to-Cell Transmission of a High-Titer Hepatitis C Virus Genotype 5a Core-NS2 JFH1-Based Recombinant.J Virol. 2015 Aug;89(15):7758-75. doi: 10.1128/JVI.00039-15. Epub 2015 May 20. J Virol. 2015. PMID: 25995244 Free PMC article.
-
Patch-Clamp Study of Hepatitis C p7 Channels Reveals Genotype-Specific Sensitivity to Inhibitors.Biophys J. 2016 Jun 7;110(11):2419-2429. doi: 10.1016/j.bpj.2016.04.018. Biophys J. 2016. PMID: 27276260 Free PMC article.
-
Antibodies Targeting Novel Neutralizing Epitopes of Hepatitis C Virus Glycoprotein Preclude Genotype 2 Virus Infection.PLoS One. 2015 Sep 25;10(9):e0138756. doi: 10.1371/journal.pone.0138756. eCollection 2015. PLoS One. 2015. PMID: 26406225 Free PMC article.
-
Human Immunodeficiency Viruses Pseudotyped with SARS-CoV-2 Spike Proteins Infect a Broad Spectrum of Human Cell Lines through Multiple Entry Mechanisms.Viruses. 2021 May 21;13(6):953. doi: 10.3390/v13060953. Viruses. 2021. PMID: 34064066 Free PMC article.
-
Antigenic and immunogenic evaluation of permutations of soluble hepatitis C virus envelope protein E2 and E1 antigens.PLoS One. 2021 Jul 30;16(7):e0255336. doi: 10.1371/journal.pone.0255336. eCollection 2021. PLoS One. 2021. PMID: 34329365 Free PMC article.
References
-
- Fauvelle C, Lepiller Q, Felmlee DJ, Fofana I, Habersetzer F, Stoll-Keller F, et al. Hepatitis C virus vaccines – Progress and perspectives. Microb. Pathog. 2013;58:66–72. - PubMed
-
- Alter HJ, Seeff LB. Recovery, Persistence, and Sequelae in Hepatitis C Virus Infection: A Perspective on Long-Term Outcome. Semin. Liver Dis. 2000;20:0017–0036. - PubMed
-
- Simmonds P, Bukh J, Combet C, Deléage G, Enomoto N, Feinstone S, et al. Consensus proposals for a unified system of nomenclature of hepatitis C virus genotypes. Hepatology. 2005;42:962–973. - PubMed
-
- Gottwein JM, Bukh J. Chapter 2 Cutting the Gordian Knot-Development and Biological Relevance of Hepatitis C Virus Cell Culture Systems. Adv. Virus Res. 2008;71:51–133. - PubMed
-
- Amoroso P, Rapicetta M, Tosti ME, Mele A, Spada E, Buonocore S, et al. Correlation between virus genotype and chronicity rate in acute hepatitis C. J. Hepatol. 1998;28:939–944. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

