Interactive XCMS Online: simplifying advanced metabolomic data processing and subsequent statistical analyses
- PMID: 24934772
- PMCID: PMC4215863
- DOI: 10.1021/ac500734c
Interactive XCMS Online: simplifying advanced metabolomic data processing and subsequent statistical analyses
Abstract
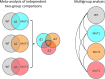
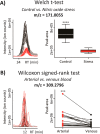
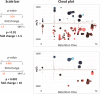
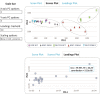
XCMS Online (xcmsonline.scripps.edu) is a cloud-based informatic platform designed to process and visualize mass-spectrometry-based, untargeted metabolomic data. Initially, the platform was developed for two-group comparisons to match the independent, "control" versus "disease" experimental design. Here, we introduce an enhanced XCMS Online interface that enables users to perform dependent (paired) two-group comparisons, meta-analysis, and multigroup comparisons, with comprehensive statistical output and interactive visualization tools. Newly incorporated statistical tests cover a wide array of univariate analyses. Multigroup comparison allows for the identification of differentially expressed metabolite features across multiple classes of data while higher order meta-analysis facilitates the identification of shared metabolic patterns across multiple two-group comparisons. Given the complexity of these data sets, we have developed an interactive platform where users can monitor the statistical output of univariate (cloud plots) and multivariate (PCA plots) data analysis in real time by adjusting the threshold and range of various parameters. On the interactive cloud plot, metabolite features can be filtered out by their significance level (p-value), fold change, mass-to-charge ratio, retention time, and intensity. The variation pattern of each feature can be visualized on both extracted-ion chromatograms and box plots. The interactive principal component analysis includes scores, loadings, and scree plots that can be adjusted depending on scaling criteria. The utility of XCMS functionalities is demonstrated through the metabolomic analysis of bacterial stress response and the comparison of lymphoblastic leukemia cell lines.
Figures



References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

