The specification of imprints in mammals
- PMID: 24939713
- PMCID: PMC4105455
- DOI: 10.1038/hdy.2014.54
The specification of imprints in mammals
Abstract
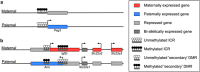
At the heart of genomic imprinting in mammals are imprinting control regions (ICRs), which are the discrete genetic elements that confer imprinted monoallelic expression to several genes in imprinted gene clusters. A characteristic of the known ICRs is that they acquire different epigenetic states, exemplified by differences in DNA methylation, in the sperm and egg, and these imprint marks remain on the sperm- and oocyte-derived alleles into the next generation as a lifelong memory of parental origin. Although there has been much focus on gametic marking of ICRs as the point of imprint specification, recent mechanistic studies and genome-wide DNA methylation profiling do not support the existence of a specific imprinting machinery in germ cells. Rather, ICRs are part of more widespread methylation events that occur during gametogenesis. Instead, a decisive component in the specification of imprints is the choice of which sites of gamete-derived methylation to maintain in the zygote and preimplantation embryo at a time when much of the remainder of the genome is being demethylated. Among the factors involved in this selection, the zinc-finger protein Zfp57 can be regarded as an imprint-specific, sequence-specific DNA binding factor responsible for maintaining methylation at most ICRs. The recent insights into the balance of gametic and zygotic contributions to imprint specification should help understand mechanistic opportunities and constraints on the evolution of imprinting in mammals.
Figures
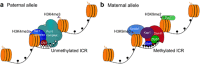
References
-
- Beaujean N, Hartshorne G, Cavilla J, Taylor J, Gardner J, Wilmut I, et al. Non-conservation of mammalian preimplantation methylation dynamics. Curr Biol. 2004;14:R266–R267. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

