A Spinach molecular beacon triggered by strand displacement
- PMID: 24942625
- PMCID: PMC4105745
- DOI: 10.1261/rna.045047.114
A Spinach molecular beacon triggered by strand displacement
Abstract
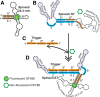
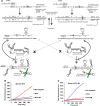
We have re-engineered the fluorescent RNA aptamer Spinach to be activated in a sequence-dependent manner. The original Spinach aptamer was extended at its 5'- and 3'-ends to create Spinach.ST, which is predicted to fold into an inactive conformation and thus prevent association with the small molecule fluorophore DFHBI. Hybridization of a specific trigger oligonucleotide to a designed toehold leads to toehold-initiated strand displacement and refolds Spinach into the active, fluorophore-binding conformation. Spinach.ST not only specifically detects its target oligonucleotide but can discriminate readily against single-nucleotide mismatches. RNA amplicons produced during nucleic acid sequence-based amplification (NASBA) of DNA or RNA targets could be specifically detected and reported in real-time by conformational activation of Spinach.ST generated by in vitro transcription. In order to adapt any target sequence to detection by a Spinach reporter we used a primer design technique that brings together otherwise distal toehold sequences via hairpin formation. The same techniques could potentially be used to adapt common Spinach reporters to non-nucleic acid analytes, rather than by making fusions between aptamers and Spinach.
Keywords: aptamer; aptamer beacon; fluorescent RNA; molecular beacon; nucleic acid diagnostics; nucleic acid engineering; strand displacement.
© 2014 Bhadra and Ellington; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures


References
-
- Andersen ES, Dong M, Nielsen MM, Jahn K, Subramani R, Mamdouh W, Golas MM, Sander B, Stark H, Oliveira CLP, et al.2009. Self-assembly of a nanoscale DNA box with a controllable lid. Nature 459: 73–76 - PubMed
-
- Babendure JR, Adams SR, Tsien RY 2003. Aptamers switch on fluorescence of triphenylmethane dyes. J Am Chem Soc 125: 14716–14717 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
