Improved sensitivity for molecular detection of bacterial and Candida infections in blood
- PMID: 24951806
- PMCID: PMC4313132
- DOI: 10.1128/JCM.00801-14
Improved sensitivity for molecular detection of bacterial and Candida infections in blood
Abstract
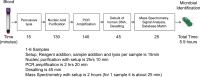
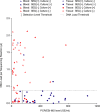
The rapid identification of bacteria and fungi directly from the blood of patients with suspected bloodstream infections aids in diagnosis and guides treatment decisions. The development of an automated, rapid, and sensitive molecular technology capable of detecting the diverse agents of such infections at low titers has been challenging, due in part to the high background of genomic DNA in blood. PCR followed by electrospray ionization mass spectrometry (PCR/ESI-MS) allows for the rapid and accurate identification of microorganisms but with a sensitivity of about 50% compared to that of culture when using 1-ml whole-blood specimens. Here, we describe a new integrated specimen preparation technology that substantially improves the sensitivity of PCR/ESI-MS analysis. An efficient lysis method and automated DNA purification system were designed for processing 5 ml of whole blood. In addition, PCR amplification formulations were optimized to tolerate high levels of human DNA. An analysis of 331 specimens collected from patients with suspected bloodstream infections resulted in 35 PCR/ESI-MS-positive specimens (10.6%) compared to 18 positive by culture (5.4%). PCR/ESI-MS was 83% sensitive and 94% specific compared to culture. Replicate PCR/ESI-MS testing from a second aliquot of the PCR/ESI-MS-positive/culture-negative specimens corroborated the initial findings in most cases, resulting in increased sensitivity (91%) and specificity (99%) when confirmed detections were considered true positives. The integrated solution described here has the potential to provide rapid detection and identification of organisms responsible for bloodstream infections.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures



References
-
- Kumar A, Roberts D, Wood KE, Light B, Parrillo JE, Sharma S, Suppes R, Feinstein D, Zanotti S, Taiberg L, Gurka D, Kumar A, Cheang M. 2006. Duration of hypotension before initiation of effective antimicrobial therapy is the critical determinant of survival in human septic shock. Crit. Care Med. 34:1589–1596. 10.1097/01.CCM.0000217961.75225.E9. - DOI - PubMed
-
- Dellinger RP, Levy MM, Rhodes A, Annane D, Gerlach H, Opal SM, Sevransky JE, Sprung CL, Douglas IS, Jaeschke R, Osborn TM, Nunnally ME, Townsend SR, Reinhart K, Kleinpell RM, Angus DC, Deutschman CS, Machado FR, Rubenfeld GD, Webb SA, Beale RJ, Vincent JL, Moreno R, Surviving Sepsis Campaign Guidelines Committee including the Pediatric Subgroup 2013. Surviving Sepsis Campaign: international guidelines for management of severe sepsis and septic shock: 2012. Crit. Care Med. 41:580–637. 10.1097/CCM.0b013e31827e83af. - DOI - PubMed
-
- Dellinger RP, Levy MM, Carlet JM, Bion J, Parker MM, Jaeschke R, Reinhart K, Angus DC, Brun-Buisson C, Beale R, Calandra T, Dhainaut JF, Gerlach H, Harvey M, Marini JJ, Marshall J, Ranieri M, Ramsay G, Sevransky J, Thompson BT, Townsend S, Vender JS, Zimmerman JL, Vincent JL, International Surviving Sepsis Campaign Guidelines Committee, American Association of Critical-Care Nurses, American College of Chest Physicians, American College of Emergency Physicians, Canadian Critical Care Society, European Society of Clinical Microbiology and Infectious Diseases, European Society of Intensive Care Medicine, European Respiratory Society, International Sepsis Forum, Japanese Association for Acute Medicine, Japanese Society of Intensive Care Medicine, Society of Critical Care Medicine, Society of Hospital Medicine, Surgical Infection Society, World Federation of Societies of Intensive and Critical Care Medicine 2008. Surviving Sepsis Campaign: international guidelines for management of severe sepsis and septic shock: 2008. Crit. Care Med. 36:296–327. 10.1097/01.CCM.0000298158.12101.41. - DOI - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

