Regulation of histone methylation by noncoding RNAs
- PMID: 24954181
- PMCID: PMC4258512
- DOI: 10.1016/j.bbagrm.2014.06.006
Regulation of histone methylation by noncoding RNAs
Abstract
Cells can adapt to their environment and develop distinct identities by rewiring their transcriptional networks to regulate the output of key biological pathways without concomitant mutations to the underlying genes. These alterations, called epigenetic changes, persist stably through mitotic or, in some instances, meiotic cell divisions. In eukaryotes, heritable changes to chromatin structure are a prominent, but not exclusive, mechanism by which epigenetic changes are mediated. These changes are initiated by sequence-specific events, which trigger a cascade of molecular interactions resulting in feedback mechanisms, alterations in chromatin structure, histone posttranslational modifications (PTMs), and ultimately establishment of distinct transcriptional states. In recent years, advances in next generation sequencing have led to the discovery of several novel classes of noncoding RNAs (ncRNAs). In addition to their well-established cytoplasmic roles in posttranscriptional regulation of gene expression, ncRNAs have emerged as key regulators of epigenetic changes via chromatin-dependent mechanisms in organisms ranging from yeast to man. They function by affecting chromatin structure, histone PTMs, and the recruitment of transcriptional activating or repressing complexes. Among histone PTMs, lysine methylation serves as the binding substrate for the recruitment of key protein complexes involved in the regulation of genome architecture, stability, and gene expression. In this review, we will outline the known mechanisms by which ncRNAs of different origins regulate histone methylation, and in doing so contribute to a variety of genome regulatory functions in eukaryotes.
Keywords: Epigenetics; Histone methylation; Noncoding RNA; lncRNA; piRNA; siRNA.
Copyright © 2014 Elsevier B.V. All rights reserved.
Figures
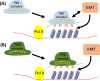
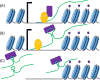
Similar articles
-
Signals and combinatorial functions of histone modifications.Annu Rev Biochem. 2011;80:473-99. doi: 10.1146/annurev-biochem-061809-175347. Annu Rev Biochem. 2011. PMID: 21529160 Review.
-
Epigenetics of Huntington's Disease.Adv Exp Med Biol. 2017;978:277-299. doi: 10.1007/978-3-319-53889-1_15. Adv Exp Med Biol. 2017. PMID: 28523552 Review.
-
[Advances in histone methyltransferases and histone demethylases].Ai Zheng. 2008 Oct;27(10):1018-25. Ai Zheng. 2008. PMID: 18851779 Review. Chinese.
-
Histone lysine methylation and chromatin replication.Biochim Biophys Acta. 2014 Dec;1839(12):1433-9. doi: 10.1016/j.bbagrm.2014.03.009. Epub 2014 Mar 28. Biochim Biophys Acta. 2014. PMID: 24686120 Review.
-
Histone H4 lysine 20 methylation: key player in epigenetic regulation of genomic integrity.Nucleic Acids Res. 2013 Mar 1;41(5):2797-806. doi: 10.1093/nar/gkt012. Epub 2013 Jan 23. Nucleic Acids Res. 2013. PMID: 23345616 Free PMC article. Review.
Cited by
-
Obesity II: Establishing causal links between chemical exposures and obesity.Biochem Pharmacol. 2022 May;199:115015. doi: 10.1016/j.bcp.2022.115015. Epub 2022 Apr 5. Biochem Pharmacol. 2022. PMID: 35395240 Free PMC article. Review.
-
PARTICLE triplexes cluster in the tumor suppressor WWOX and may extend throughout the human genome.Sci Rep. 2017 Aug 2;7(1):7163. doi: 10.1038/s41598-017-07295-5. Sci Rep. 2017. PMID: 28769061 Free PMC article.
-
Noncoding RNAs in Nonalcoholic Fatty Liver Disease: Potential Diagnosis and Prognosis Biomarkers.Dis Markers. 2020 Aug 27;2020:8822859. doi: 10.1155/2020/8822859. eCollection 2020. Dis Markers. 2020. PMID: 33133304 Free PMC article. Review.
-
Non-coding RNAs: Epigenetic regulators of bone development and homeostasis.Bone. 2015 Dec;81:746-756. doi: 10.1016/j.bone.2015.05.026. Epub 2015 May 31. Bone. 2015. PMID: 26039869 Free PMC article. Review.
-
From Environmental Epigenetics to the Inheritance of Acquired Traits: A Historian and Molecular Perspective on an Unnecessary Lamarckian Explanation.Biomolecules. 2023 Jul 5;13(7):1077. doi: 10.3390/biom13071077. Biomolecules. 2023. PMID: 37509113 Free PMC article. Review.
References
-
- Surani MA, Hayashi K, Hajkova P. Genetic and epigenetic regulators of pluripotency. Cell. 2007 Feb;128(4):747–62. - PubMed
-
- Murray AW, Szostak JW. Chromosome segregation in mitosis and meiosis. Annu Rev Cell Biol. 1985 Jan;1:289–315. - PubMed
-
- Marians KJ. Prokaryotic DNA replication. Annu Rev Biochem. 1992 Jan;61:673–719. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

