Changes in granulosa cells gene expression associated with growth, plateau and atretic phases in medium bovine follicles
- PMID: 24955130
- PMCID: PMC4046060
- DOI: 10.1186/1757-2215-7-50
Changes in granulosa cells gene expression associated with growth, plateau and atretic phases in medium bovine follicles
Abstract
Background: The objective of this study was to build the transcriptomic profile of granulosa cells originating from follicles 6 to 9 mm in diameter in dairy cattle using microarrays.
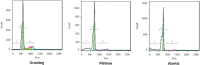
Methods: Granulosa cells originating from three different phases of antral follicle growth were compared: growing (G), plateau (P) and atresia (A), as categorized by flow cytometry profiles of DNA. The growing and atretic conditions were each hybridized against the plateau condition as a reference in order to understand the specific biological mechanisms modulated in this class of follicles.
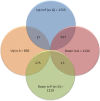
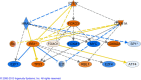
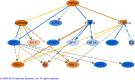
Results: 2,942 genes were differentially expressed (P < 0.05) in P vs. G and 1,974 in A vs. P. A clear segregation of the 3 phases was confirmed by between group analysis (BGA). The first characteristic of the plateau phase is the activation of the upstream regulators TP53 and PTEN which participate in the reduction of cell growth through MYC, FOS and E2F1-2-3. We also observed the down-regulation of steroidogenesis genes: CYP11A1 and CYP19A1, in the granulosa cells of the plateau phase relative to the growth phase. On the other hand, the A vs. P contrast showed up-regulation of multiple transcripts associated to apoptosis: CCT2, DAB2, DSG2 and TGM2.
Conclusions: This study offers multiple candidate genes to be further studied in order to elucidate their role in the modulation of follicular development and, ultimately, of oocyte quality.
Keywords: Cow; Granulosa cells; Growth status; Transcriptome.
Figures



References
-
- Erickson BH. Development and senescence of the postnatal bovine ovary. J Anim Sci. 1966;25(3):800–805. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

