Metabolomic and Lipidomic Analysis of the Heart of Peroxisome Proliferator-Activated Receptor-γ Coactivator 1-β Knock Out Mice on a High Fat Diet
- PMID: 24957515
- PMCID: PMC3901207
- DOI: 10.3390/metabo2020366
Metabolomic and Lipidomic Analysis of the Heart of Peroxisome Proliferator-Activated Receptor-γ Coactivator 1-β Knock Out Mice on a High Fat Diet
Abstract
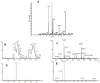
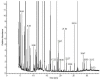
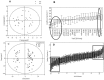
The peroxisome proliferator-activated receptor-γ coactivators (PGC-1) are transcriptional coactivators with an important role in mitochondrial biogenesis and regulation of genes involved in the electron transport chain and oxidative phosphorylation in oxidative tissues including cardiac tissue. These coactivators are thought to play a key role in the development of obesity, type 2 diabetes and the metabolic syndrome. In this study we have used a combined metabolomic and lipidomic analysis of cardiac tissue from the PGC-1β null mouse to examine the effects of a high fat diet on this organ. Multivariate statistics readily separated tissue from PGC-1β null mice from their wild type controls either in gender specific models or in combined datasets. This was associated with an increase in creatine and a decrease in taurine in the null mouse, and an increase in myristic acid and a reduction in long chain polyunsaturated fatty acids for both genders. The most profound changes were detected by liquid chromatography mass spectrometry analysis of intact lipids with the tissue from the null mouse having a profound increase in a number of triglycerides. The metabolomic and lipodomic changes indicate PGC-1β has a profound influence on cardiac metabolism.
Figures

Similar articles
-
Rosiglitazone-induced mitochondrial biogenesis in white adipose tissue is independent of peroxisome proliferator-activated receptor γ coactivator-1α.PLoS One. 2011;6(11):e26989. doi: 10.1371/journal.pone.0026989. Epub 2011 Nov 7. PLoS One. 2011. PMID: 22087241 Free PMC article.
-
A role for peroxisome proliferator-activated receptor γ coactivator-1 in the control of mitochondrial dynamics during postnatal cardiac growth.Circ Res. 2014 Feb 14;114(4):626-36. doi: 10.1161/CIRCRESAHA.114.302562. Epub 2013 Dec 23. Circ Res. 2014. PMID: 24366168 Free PMC article.
-
The diabetes medication canagliflozin promotes mitochondrial remodelling of adipocyte via the AMPK-Sirt1-Pgc-1α signalling pathway.Adipocyte. 2020 Dec;9(1):484-494. doi: 10.1080/21623945.2020.1807850. Adipocyte. 2020. PMID: 32835596 Free PMC article.
-
PGC-1beta: a co-activator that sets the tone for both basal and stress-stimulated mitochondrial activity.Adv Exp Med Biol. 2009;646:133-9. doi: 10.1007/978-1-4020-9173-5_15. Adv Exp Med Biol. 2009. PMID: 19536672 Review.
-
Natural products, PGC-1 α , and Duchenne muscular dystrophy.Acta Pharm Sin B. 2020 May;10(5):734-745. doi: 10.1016/j.apsb.2020.01.001. Epub 2020 Jan 8. Acta Pharm Sin B. 2020. PMID: 32528825 Free PMC article. Review.
Cited by
-
Evaluation of extraction protocols for simultaneous polar and non-polar yeast metabolite analysis using multivariate projection methods.Metabolites. 2013 Jul 23;3(3):592-605. doi: 10.3390/metabo3030592. Metabolites. 2013. PMID: 24958140 Free PMC article.
-
The Cardiac Lipidome in Models of Cardiovascular Disease.Metabolites. 2020 Jun 17;10(6):254. doi: 10.3390/metabo10060254. Metabolites. 2020. PMID: 32560541 Free PMC article. Review.
References
-
- Scheuermann-Freestone M., Madsen P.L., Manners D., Blamire A.M., Buckingham R.E., Styles P., Radda G.K., Neubauer S., Clarke K. Abnormal cardiac and skeletal muscle energy metabolism in patients with type 2 diabetes. Circulation. 2003;107:3040–3046. - PubMed
-
- King L.M., Sidell R.J., Wilding J.R., Radda G.K., Clarke K. Free fatty acids, but not ketone bodies, protect diabetic rat hearts during low-flow ischemia. Am. J. Physiol., Heart Circ. Physiol. 2001;280:H1173–H1181. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

