Metabolic Consequences of TGFb Stimulation in CulturedPrimary Mouse Hepatocytes Screened from Transcript Data with ModeScore
- PMID: 24957771
- PMCID: PMC3901234
- DOI: 10.3390/metabo2040983
Metabolic Consequences of TGFb Stimulation in CulturedPrimary Mouse Hepatocytes Screened from Transcript Data with ModeScore
Abstract
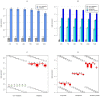
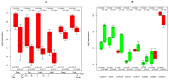
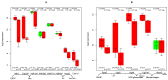
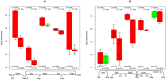
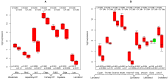
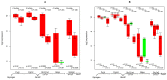
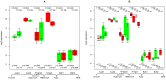
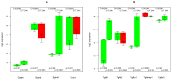
TGFb signaling plays a major role in the reorganization of liver tissue upon injury and is an important driver of chronic liver disease. This is achieved by a deep impact on a cohort of cellular functions. To comprehensively assess the full range of affected metabolic functions, transcript changes of cultured mouse hepatocytes were analyzed with a novel method (ModeScore), which predicts the activity of metabolic functions by scoring transcript expression changes with 987 reference flux distributions, which yielded the following hypotheses. TGFb multiplies down-regulation of most metabolic functions occurring in culture stressed controls. This is especially pronounced for tyrosine degradation, urea synthesis, glucuronization capacity, and cholesterol synthesis. Ethanol degradation and creatine synthesis are down-regulated only in TGFb treated hepatocytes, but not in the control. Among the few TGFb dependently up-regulated functions, synthesis of various collagens is most pronounced. Further interesting findings include: down-regulation of glucose export is postponed by TGFb, TGFb up-regulates the synthesis capacity of ketone bodies only as an early response, TGFb suppresses the strong up-regulation of Vanin, and TGFb induces re-formation of ceramides and sphingomyelin.
Figures
References
-
- Liska V., Treska V., Mirka H., Kobr J., Sykora R., Skalicky T., Sutnar A., Vycital O., Bruha J., Pitule P., et al. Inhibition of transforming growth factor beta-1 augments liver regeneration after partial portal vein ligation in a porcine experimental model. Hepatogastroenterology. 2012;59:235–240. - PubMed
-
- Klingmüller U., Bauer A., Bohl S., Nickel P.J., Breitkopf K., Dooley S., Zellmer S., Kern C., Merfort I., Sparna T., et al. Primary mouse hepatocytes for systems biology approaches: A standardized in vitro system for modelling of signal transduction pathways. Syst. Biol. 2006;153:433–447. doi: 10.1049/ip-syb:20050067. - DOI - PubMed
-
- Zeilinger K., Auth S., Unger J., Grebe A., Mao L., Petrik M., Holland G., Appel K., Nüssler A., Neuhaus P., et al. Liver cell culture in bioreactors for in vitro drug studies as an alternative to animal testing. ALTEX. 2000;17:3–10. - PubMed
LinkOut - more resources
Full Text Sources

