GABP transcription factor (nuclear respiratory factor 2) is required for mitochondrial biogenesis
- PMID: 24958105
- PMCID: PMC4135556
- DOI: 10.1128/MCB.00492-12
GABP transcription factor (nuclear respiratory factor 2) is required for mitochondrial biogenesis
Abstract
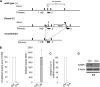
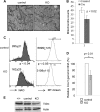
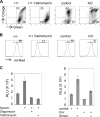
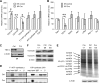
Mitochondria are membrane-bound cytoplasmic organelles that serve as the major source of ATP production in eukaryotic cells. GABP (also known as nuclear respiratory factor 2) is a nuclear E26 transformation-specific transcription factor (ETS) that binds and activates mitochondrial genes that are required for electron transport and oxidative phosphorylation. We conditionally deleted Gabpa, the DNA-binding component of this transcription factor complex, from mouse embryonic fibroblasts (MEFs) to examine the role of Gabp in mitochondrial biogenesis, function, and gene expression. Gabpα loss modestly reduced mitochondrial mass, ATP production, oxygen consumption, and mitochondrial protein synthesis but did not alter mitochondrial morphology, membrane potential, apoptosis, or the expression of several genes that were previously reported to be GABP targets. However, the expression of Tfb1m, a methyltransferase that modifies ribosomal rRNA and is required for mitochondrial protein translation, was markedly reduced in Gabpα-null MEFs. We conclude that Gabp regulates Tfb1m expression and plays an essential, nonredundant role in mitochondrial biogenesis.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures


Similar articles
-
GABP transcription factor is required for myeloid differentiation, in part, through its control of Gfi-1 expression.Blood. 2011 Aug 25;118(8):2243-53. doi: 10.1182/blood-2010-07-298802. Epub 2011 Jun 24. Blood. 2011. PMID: 21705494 Free PMC article.
-
Regulation of nephrin gene by the Ets transcription factor, GA-binding protein.Nephrol Dial Transplant. 2013 Apr;28(4):846-55. doi: 10.1093/ndt/gfs482. Epub 2012 Nov 29. Nephrol Dial Transplant. 2013. PMID: 23197680
-
Expression of the human TIMM23 and TIMM23B genes is regulated by the GABP transcription factor.Biochim Biophys Acta Gene Regul Mech. 2018 Feb;1861(2):80-94. doi: 10.1016/j.bbagrm.2018.01.006. Epub 2018 Feb 3. Biochim Biophys Acta Gene Regul Mech. 2018. PMID: 29413900
-
GA-binding protein transcription factor: a review of GABP as an integrator of intracellular signaling and protein-protein interactions.Blood Cells Mol Dis. 2004 Jan-Feb;32(1):143-54. doi: 10.1016/j.bcmd.2003.09.005. Blood Cells Mol Dis. 2004. PMID: 14757430 Review.
-
Structural organization and transcription regulation of nuclear genes encoding the mammalian cytochrome c oxidase complex.Prog Nucleic Acid Res Mol Biol. 1998;61:309-44. doi: 10.1016/s0079-6603(08)60830-2. Prog Nucleic Acid Res Mol Biol. 1998. PMID: 9752724 Review.
Cited by
-
Opposite phenotypes of muscle strength and locomotor function in mouse models of partial trisomy and monosomy 21 for the proximal Hspa13-App region.PLoS Genet. 2015 Mar 24;11(3):e1005062. doi: 10.1371/journal.pgen.1005062. eCollection 2015 Mar. PLoS Genet. 2015. PMID: 25803843 Free PMC article.
-
Evidence for ~12-h ultradian gene programs in humans.NPJ Biol Timing Sleep. 2024;1(1):4. doi: 10.1038/s44323-024-00005-1. Epub 2024 Aug 9. NPJ Biol Timing Sleep. 2024. PMID: 39148626 Free PMC article.
-
EPA and DHA Enhance CACT Promoter Activity by GABP/NRF2.Int J Mol Sci. 2024 Aug 22;25(16):9095. doi: 10.3390/ijms25169095. Int J Mol Sci. 2024. PMID: 39201781 Free PMC article.
-
How an increase in the copy number of HSV-1 during latency can cause Alzheimer's disease: the viral and cellular dynamics according to the microcompetition model.J Neurovirol. 2021 Dec;27(6):895-916. doi: 10.1007/s13365-021-01012-9. Epub 2021 Oct 11. J Neurovirol. 2021. PMID: 34635992
-
Identification of GA-Binding Protein Transcription Factor Alpha Subunit (GABPA) as a Novel Bookmarking Factor.Int J Mol Sci. 2019 Mar 4;20(5):1093. doi: 10.3390/ijms20051093. Int J Mol Sci. 2019. PMID: 30836589 Free PMC article.
References
-
- Litonin D, Sologub M, Shi Y, Savkina M, Anikin M, Falkenberg M, Gustafsson CM, Temiakov D. 2010. Human mitochondrial transcription revisited: only TFAM and TFB2M are required for transcription of the mitochondrial genes in vitro. J. Biol. Chem. 285:18129–18133. 10.1074/jbc.C110.128918 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
