Protective role of the capsule and impact of serotype 4 switching on Streptococcus mitis
- PMID: 24958712
- PMCID: PMC4187822
- DOI: 10.1128/IAI.01840-14
Protective role of the capsule and impact of serotype 4 switching on Streptococcus mitis
Abstract
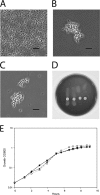
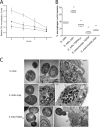
The polysaccharide capsule surrounding Streptococcus pneumoniae is essential for virulence. Recently, Streptococcus mitis, a human commensal and a close relative of S. pneumoniae, was also shown to have a capsule. In this study, the S. mitis type strain switched capsule by acquisition of the serotype 4 capsule locus of S. pneumoniae TIGR4, following induction of competence for natural transformation. Comparison of the wild type with the capsule-switching mutant and with a capsule deletion mutant showed that the capsule protected S. mitis against phagocytosis by RAW 264.7 macrophages. This effect was enhanced in the S. mitis strain expressing the S. pneumoniae capsule, which showed, in addition, increased resistance against early clearance in a mouse model of lung infection. Expression of both capsules also favored survival in human blood, and the effect was again more pronounced for the capsule-switching mutant. S. mitis survival in horse blood or in a mouse model of bacteremia was not significantly different between the wild type and the mutant strains. In all models, S. pneumoniae TIGR4 showed higher rates of survival than the S. mitis type strain or the capsule-switching mutant, except in the lung model, in which significant differences between S. pneumoniae TIGR4 and the capsule-switching mutant were not observed. Thus, we identified conditions that showed a protective function for the capsule in S. mitis. Under such conditions, S. mitis resistance to clearance could be enhanced by capsule switching to serotype 4, but it was enhanced to levels lower than those for the virulent strain S. pneumoniae TIGR4.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures


References
-
- Bentley SD, Aanensen DM, Mavroidi A, Saunders D, Rabbinowitsch E, Collins M, Donohoe K, Harris D, Murphy L, Quail MA, Samuel G, Skovsted IC, Kaltoft MS, Barrell B, Reeves PR, Parkhill J, Spratt BG. 2006. Genetic analysis of the capsular biosynthetic locus from all 90 pneumococcal serotypes. PLoS Genet. 2:262–269. 10.1371/journal.pgen.0020031 - DOI - PMC - PubMed
-
- Croucher NJ, Harris SR, Fraser C, Quail MA, Burton J, van der Linden M, McGee L, von Gottberg A, Song JH, Ko KS, Pichon B, Baker S, Parry CM, Lambertsen LM, Shahinas D, Pillai DR, Mitchell TJ, Dougan G, Tomasz A, Klugman KP, Parkhill J, Hanage WP, Bentley SD. 2011. Rapid pneumococcal evolution in response to clinical interventions. Science 331:430–434. 10.1126/science.1198545 - DOI - PMC - PubMed
-
- Golubchik T, Brueggemann AB, Street T, Gertz RE, Jr, Spencer CC, Ho T, Giannoulatou E, Link-Gelles R, Harding RM, Beall B, Peto TE, Moore MR, Donnelly P, Crook DW, Bowden R. 2012. Pneumococcal genome sequencing tracks a vaccine escape variant formed through a multi-fragment recombination event. Nat. Genet. 44:352–355. 10.1038/ng.1072 - DOI - PMC - PubMed
-
- Wyres KL, Lambertsen LM, Croucher NJ, McGee L, von Gottberg A, Linares J, Jacobs MR, Kristinsson KG, Beall BW, Klugman KP, Parkhill J, Hakenbeck R, Bentley SD, Brueggemann AB. 2013. Pneumococcal capsular switching: a historical perspective. J. Infect. Dis. 207:439–449. 10.1093/infdis/jis703 - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

