Novel components of the flagellar system in epsilonproteobacteria
- PMID: 24961693
- PMCID: PMC4073491
- DOI: 10.1128/mBio.01349-14
Novel components of the flagellar system in epsilonproteobacteria
Abstract
Motility is essential for the pathogenesis of many bacterial species. Most bacteria move using flagella, which are multiprotein filaments that rotate propelled by a cell wall-anchored motor using chemical energy. Although some components of the flagellar apparatus are common to many bacterial species, recent studies have shown significant differences in the flagellar structures of different bacterial species. The molecular bases for these differences, however, are not understood. The flagella from epsilonproteobacteria, which include the bacterial pathogens Campylobacter jejuni and Helicobacter pylori, are among the most divergent. Using next-generation sequencing combined with transposon mutagenesis, we have conducted a comprehensive high-throughput genetic screen in Campylobacter jejuni, which identified several novel components of its flagellar system. Biochemical analyses detected interactions between the identified proteins and known components of the flagellar machinery, and in vivo imaging located them to the bacterial poles, where flagella assemble. Most of the identified new components are conserved within but restricted to epsilonproteobacteria. These studies provide insight into the divergent flagella of this group of bacteria and highlight the complexity of this remarkable structure, which has adapted to carry out its conserved functions in the context of widely diverse bacterial species.
Importance: Motility is essential for the normal physiology and pathogenesis of many bacterial species. Most bacteria move using flagella, which are multiprotein filaments that rotate propelled by a motor that uses chemical energy as fuel. Although some components of the flagellar apparatus are common to many bacterial species, recent studies have shown significant divergence in the flagellar structures across bacterial species. However, the molecular bases for these differences are not understood. The flagella from epsilonproteobacteria, which include the bacterial pathogens Campylobacter jejuni and Helicobacter pylori, are among the most divergent. We conducted a comprehensive genetic screen in Campylobacter jejuni and identified several novel components of the flagellar system. These studies provide important information to understand how flagella have adapted to function in the context of widely diverse sets of bacterial species and bring unique insight into the evolution and function of this remarkable bacterial organelle.
Copyright © 2014 Gao et al.
Figures





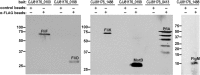


Similar articles
-
A Chaperone for the Stator Units of a Bacterial Flagellum.mBio. 2019 Aug 6;10(4):e01732-19. doi: 10.1128/mBio.01732-19. mBio. 2019. PMID: 31387912 Free PMC article.
-
Polar flagellar biosynthesis and a regulator of flagellar number influence spatial parameters of cell division in Campylobacter jejuni.PLoS Pathog. 2011 Dec;7(12):e1002420. doi: 10.1371/journal.ppat.1002420. Epub 2011 Dec 1. PLoS Pathog. 2011. PMID: 22144902 Free PMC article.
-
ChePep controls Helicobacter pylori Infection of the gastric glands and chemotaxis in the Epsilonproteobacteria.mBio. 2011 Jul 26;2(4):e00098-11. doi: 10.1128/mBio.00098-11. Print 2011. mBio. 2011. PMID: 21791582 Free PMC article.
-
Change is good: variations in common biological mechanisms in the epsilonproteobacterial genera Campylobacter and Helicobacter.Microbiol Mol Biol Rev. 2011 Mar;75(1):84-132. doi: 10.1128/MMBR.00035-10. Microbiol Mol Biol Rev. 2011. PMID: 21372321 Free PMC article. Review.
-
Motility and chemotaxis in Campylobacter and Helicobacter.Annu Rev Microbiol. 2011;65:389-410. doi: 10.1146/annurev-micro-090110-102908. Annu Rev Microbiol. 2011. PMID: 21939377 Free PMC article. Review.
Cited by
-
Essential Two-Component Systems Regulating Cell Envelope Functions: Opportunities for Novel Antibiotic Therapies.J Membr Biol. 2018 Feb;251(1):75-89. doi: 10.1007/s00232-017-9995-5. Epub 2017 Nov 2. J Membr Biol. 2018. PMID: 29098331 Review.
-
FlgY, PflA, and PflB form a spoke-ring network in the high-torque flagellar motor of Helicobacter pylori.Proc Natl Acad Sci U S A. 2025 Apr 29;122(17):e2421632122. doi: 10.1073/pnas.2421632122. Epub 2025 Apr 22. Proc Natl Acad Sci U S A. 2025. PMID: 40261933
-
FlgV forms a flagellar motor ring that is required for optimal motility of Helicobacter pylori.PLoS One. 2023 Nov 17;18(11):e0287514. doi: 10.1371/journal.pone.0287514. eCollection 2023. PLoS One. 2023. PMID: 37976320 Free PMC article.
-
A Chaperone for the Stator Units of a Bacterial Flagellum.mBio. 2019 Aug 6;10(4):e01732-19. doi: 10.1128/mBio.01732-19. mBio. 2019. PMID: 31387912 Free PMC article.
-
Molecular and Cell Biological Analysis of SwrB in Bacillus subtilis.J Bacteriol. 2021 Aug 9;203(17):e0022721. doi: 10.1128/JB.00227-21. Epub 2021 Aug 9. J Bacteriol. 2021. PMID: 34124944 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

