L-carnitine and PPARα-agonist fenofibrate are involved in the regulation of Carnitine Acetyltransferase (CrAT) mRNA levels in murine liver cells
- PMID: 24962334
- PMCID: PMC4089027
- DOI: 10.1186/1471-2164-15-514
L-carnitine and PPARα-agonist fenofibrate are involved in the regulation of Carnitine Acetyltransferase (CrAT) mRNA levels in murine liver cells
Abstract
Background: The carnitine acetyltransferase (CrAT) is a mitochondrial matrix protein that directly influences intramitochondrial acetyl-CoA pools. Murine CrAT is encoded by a single gene located in the opposite orientation head to head to the PPP2R4 gene, sharing a very condensed bi-directional promoter. Since decreased CrAT expression is correlated with metabolic inflexibility and subsequent pathological consequences, our aim was to reveal and define possible activators of CrAT transcription in the normal embryonic murine liver cell line BNL CL. 2 and via which nuclear factors based on key metabolites mainly regulate hepatic expression of CrAT. Here we describe a functional characterization of the CrAT promoter region under conditions of L-carnitine deficiency and supplementation as well as fenofibrate induction in cell culture cells.
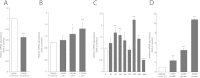
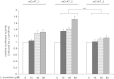
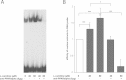
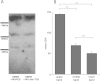
Results: The murine CrAT promoter displays some characteristics of a housekeeping gene: it lacks a TATA-box, is very GC-rich and harbors two Sp1 binding sites. Analysis of the promoter activity of CrAT by luciferase assays uncovered a L-carnitine sensitive region within -342 bp of the transcription start. Electrophoretic mobility shift and supershift assays proved the sequence element (-228/-222) to be an L-carnitine sensitive RXRα binding site, which also showed sensitivity to application of anti-PPARα and anti-PPARbp antibodies. In addition we analysed this specific RXRα/PPARα site by Southwestern Blotting technique and could pin down three protein factors binding to this promoter element. By qPCR we could quantify the nutrigenomic effect of L-carnitine itself and fenofibrate.
Conclusions: Our results indicate a cooperative interplay of L-carnitine and PPARα in transcriptional regulation of murine CrAT, which is of nutrigenomical relevance. We created experimental proof that the muCrAT gene clearly is a PPARα target. Both L-carnitine and fenofibrate are inducers of CrAT transcripts, but the important hyperlipidemic drug fenofibrate being a more potent one, as a consequence of its pharmacological interaction.
Figures


Similar articles
-
L-carnitine exerts a nutrigenomic effect via direct modulation of nuclear receptor signaling in adipocytes, hepatocytes and SKMC, demonstrating its nutritional impact.Nutr Res. 2021 Jan;85:84-98. doi: 10.1016/j.nutres.2020.11.003. Epub 2020 Nov 16. Nutr Res. 2021. PMID: 33453499
-
Regulation of Octn2 transporter (SLC22A5) by peroxisome proliferator activated receptor alpha.Biol Pharm Bull. 2008 Jun;31(6):1230-6. doi: 10.1248/bpb.31.1230. Biol Pharm Bull. 2008. PMID: 18520060
-
Redesign of carnitine acetyltransferase specificity by protein engineering.J Biol Chem. 2004 Aug 6;279(32):33899-908. doi: 10.1074/jbc.M402685200. Epub 2004 May 21. J Biol Chem. 2004. PMID: 15155769
-
l-carnitine supplementation during vitrification or warming of in vivo-produced ovine embryos does not affect embryonic survival rates, but alters CrAT and PRDX1 expression.Theriogenology. 2018 Jan 1;105:150-157. doi: 10.1016/j.theriogenology.2017.09.022. Epub 2017 Sep 21. Theriogenology. 2018. PMID: 28965027
-
Carnitine O-Acetyltransferase as a Central Player in Lipid and Branched-Chain Amino Acid Metabolism, Epigenetics, Cell Plasticity, and Organelle Function.Biomolecules. 2025 Feb 2;15(2):216. doi: 10.3390/biom15020216. Biomolecules. 2025. PMID: 40001519 Free PMC article. Review.
Cited by
-
L-Carnitine Is Involved in Hyperbaric Oxygen-Mediated Therapeutic Effects in High Fat Diet-Induced Lipid Metabolism Dysfunction.Molecules. 2020 Jan 1;25(1):176. doi: 10.3390/molecules25010176. Molecules. 2020. PMID: 31906305 Free PMC article.
-
Comprehensive review of the expanding roles of the carnitine pool in metabolic physiology: beyond fatty acid oxidation.J Transl Med. 2025 Mar 14;23(1):324. doi: 10.1186/s12967-025-06341-5. J Transl Med. 2025. PMID: 40087749 Free PMC article. Review.
References
-
- Strijbis K, Vaz F, Distel B. Enzymology of the carnitine biosynthesis pathway. IUBMB Life. 2010;62(5):357–362. - PubMed
-
- Tanphaichitr V, Broquist HP. Role of lysine and N-trimethyllysine in carnitine biosynthesis. J Biol Chem. 1973;248(6):2176–2181. - PubMed
-
- Kerner J, Hoppel C. Fatty acid import into mitochondria. Biochem Biophys Acta Mol Cell Biol Lipids. 2000;1486(1):1–17. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

