Diagnostically relevant facial gestalt information from ordinary photos
- PMID: 24963138
- PMCID: PMC4067075
- DOI: 10.7554/eLife.02020
Diagnostically relevant facial gestalt information from ordinary photos
Abstract
Craniofacial characteristics are highly informative for clinical geneticists when diagnosing genetic diseases. As a first step towards the high-throughput diagnosis of ultra-rare developmental diseases we introduce an automatic approach that implements recent developments in computer vision. This algorithm extracts phenotypic information from ordinary non-clinical photographs and, using machine learning, models human facial dysmorphisms in a multidimensional 'Clinical Face Phenotype Space'. The space locates patients in the context of known syndromes and thereby facilitates the generation of diagnostic hypotheses. Consequently, the approach will aid clinicians by greatly narrowing (by 27.6-fold) the search space of potential diagnoses for patients with suspected developmental disorders. Furthermore, this Clinical Face Phenotype Space allows the clustering of patients by phenotype even when no known syndrome diagnosis exists, thereby aiding disease identification. We demonstrate that this approach provides a novel method for inferring causative genetic variants from clinical sequencing data through functional genetic pathway comparisons.DOI: http://dx.doi.org/10.7554/eLife.02020.001.
Keywords: clinical genetics; computational biology; computer vision; phenotyping.
Copyright © 2014, Ferry et al.
Conflict of interest statement
CPP: Senior editor,
The other authors declare that no competing interests exist.
Figures





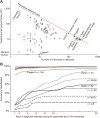
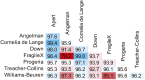
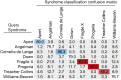
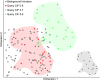
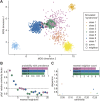
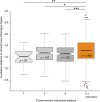

References
-
- Allanson JE, Bohring A, Dorr HG, Dufke A, Gillessen-Kaesbach G, Horn D, Konig R, Kratz CP, Kutsche K, Pauli S, Raskin S, Rauch A, Turner A, Wieczorek D, Zenker M. The face of Noonan syndrome: does phenotype predict genotype. American Journal of Medical Genetics. 2010;152A:1960–1966. doi: 10.1002/ajmg.a.33518. - DOI - PMC - PubMed
-
- Bastian M, Heymann S, Jacomy M. Gephi: An open source software for exploring and manipulating networks. AAAI Publications, Third International AAAI Conference on Weblogs and Social Media 2009
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

