The impact of first trimester phthalate and phenol exposure on IGF2/H19 genomic imprinting and birth outcomes
- PMID: 24972507
- PMCID: PMC4155603
- DOI: 10.1016/j.envres.2014.04.032
The impact of first trimester phthalate and phenol exposure on IGF2/H19 genomic imprinting and birth outcomes
Abstract
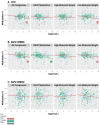
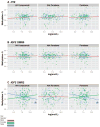
Genomic imprinting leads to parent-of-origin specific gene expression and is determined by epigenetic modification of genes. The paternally expressed gene insulin-like growth-factor 2 (IGF2) is located about ~100kb from the maternally expressed non-coding gene H19 on human chromosome 11, and both genes play major roles in embryonic and placental growth. Given adverse gestational environments can influence DNA methylation patterns in extra-embryonic tissues, we hypothesized that prenatal exposure to endocrine disrupting chemicals (EDCs) alters H19 and IGF2 methylation in placenta. Our study was restricted to a total of 196 women co-enrolled in the Predictors of Preeclampsia Study and the Harvard Epigenetic Birth Cohort. First trimester urine concentrations of 8 phenols and 11 phthalate metabolites were measured and used to characterize EDC exposure profiles. We assessed methylation of differentially methylated regions (DMRs) by pyrosequencing of H19, IGF2DMR0, and IGF2DMR2 and correlated values with phenol and phthalate metabolites. We also assessed overall expression and allele-specific expression of H19 and IGF2. We found several significant associations between DNA methylation and additive biomarker measurements. A significant decrease in H19 methylation was associated with high levels of the sum (Σ) of phthalate metabolites and metabolites of low molecular weight (LMW) phthalates. Σphthalate and LMW phthalate concentrations were inversely associated with IGF2DMR0 methylation values. Variation in methylation was not associated with changes in allele-specific expression. However increased deviation of allele-specific expression of H19 was associated with Σdi(2-ethylhexyl) phthalate metabolites and high molecular weight phthalates. Neither methylation nor expression of these imprinted regions had a significant impact on birth length or birth weight. Overall, our study provides new insight into an epigenetic mechanism that occurs following EDC exposure.
Keywords: Allele-specific expression; Endocrine disruptors; Epidemiology; Imprinting; Methylation.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures

References
-
- Barker DJ. Maternal nutrition, fetal nutrition, and disease in later life. Nutrition. 1997;13(9):807–813. - PubMed
-
- Roseboom TJ, Painter RC, van Abeelen AF, Veenendaal MV, de Rooij SR. Hungry in the womb: what are the consequences? Lessons from the Dutch famine. Maturitas. 2011;70(2):141–145. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

