Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5' sites
- PMID: 24981863
- PMCID: PMC4142486
- DOI: 10.1016/j.celrep.2014.05.048
Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5' sites
Abstract
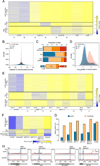
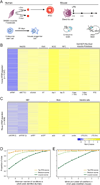
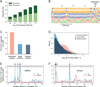
N6-methyladenosine (m6A) is a common modification of mRNA with potential roles in fine-tuning the RNA life cycle. Here, we identify a dense network of proteins interacting with METTL3, a component of the methyltransferase complex, and show that three of them (WTAP, METTL14, and KIAA1429) are required for methylation. Monitoring m6A levels upon WTAP depletion allowed the definition of accurate and near single-nucleotide resolution methylation maps and their classification into WTAP-dependent and -independent sites. WTAP-dependent sites are located at internal positions in transcripts, topologically static across a variety of systems we surveyed, and inversely correlated with mRNA stability, consistent with a role in establishing "basal" degradation rates. WTAP-independent sites form at the first transcribed base as part of the cap structure and are present at thousands of sites, forming a previously unappreciated layer of transcriptome complexity. Our data shed light on the proteomic and transcriptional underpinnings of this RNA modification.
Copyright © 2014 The Authors. Published by Elsevier Inc. All rights reserved.
Figures


References
-
- Bembom O. seqLogo: Sequence logos for DNA sequence alignments. 2011
-
- Bokar J, Rath-Shambaugh M, Ludwiczak R, Narayan P, Rottman F. Characterization and partial purification of mRNA N6-adenosine methyltransferase from HeLa cell nuclei. Internal mRNA methylation requires a multisubunit complex. The Journal of biological chemistry. 1994;269:17697–17704. - PubMed
-
- Bokar JA. The biosynthesis and functional roles of methylated nucleosides in eukaryotic mRNA. In: Grosjean Henri., editor. Fine-Tuning of RNA Functions by Modification and Editing. Vol. 12. 2005. pp. 141–177.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

