Supraspliceosomes at defined functional states portray the pre-assembled nature of the pre-mRNA processing machine in the cell nucleus
- PMID: 24983480
- PMCID: PMC4139805
- DOI: 10.3390/ijms150711637
Supraspliceosomes at defined functional states portray the pre-assembled nature of the pre-mRNA processing machine in the cell nucleus
Abstract
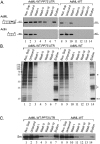
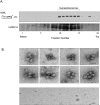
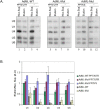
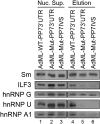
When isolated from mammalian cell nuclei, all nuclear pre-mRNAs are packaged in multi-subunit large ribonucleoprotein complexes-supraspliceosomes-composed of four native spliceosomes interconnected by the pre-mRNA. Supraspliceosomes contain all five spliceosomal U snRNPs, together with other splicing factors, and are functional in splicing. Supraspliceosomes studied thus far represent the steady-state population of nuclear pre-mRNAs that were isolated at different stages of the splicing reaction. To analyze specific splicing complexes, here, we affinity purified Pseudomonas aeruginosa phage 7 (PP7)-tagged splicing complexes assembled in vivo on Adenovirus Major Late (AdML) transcripts at specific functional stages, and characterized them using molecular techniques including mass spectrometry. First, we show that these affinity purified splicing complexes assembled on PP7-tagged AdML mRNA or on PP7-tagged AdML pre-mRNA are assembled in supraspliceosomes. Second, similar to the general population of supraspliceosomes, these defined supraspliceosomes populations are assembled with all five U snRNPs at all splicing stages. This study shows that dynamic changes in base-pairing interactions of U snRNA:U snRNA and U snRNA:pre-mRNA that occur in vivo during the splicing reaction do not require changes in U snRNP composition of the supraspliceosome. Furthermore, there is no need to reassemble a native spliceosome for the splicing of each intron, and rearrangements of the interactions will suffice.
Figures



Similar articles
-
The Supraspliceosome - A Multi-Task Machine for Regulated Pre-mRNA Processing in the Cell Nucleus.Comput Struct Biotechnol J. 2014 Sep 28;11(19):113-22. doi: 10.1016/j.csbj.2014.09.008. eCollection 2014 Sep. Comput Struct Biotechnol J. 2014. PMID: 25408845 Free PMC article. Review.
-
Structural studies of the endogenous spliceosome - The supraspliceosome.Methods. 2017 Aug 1;125:70-83. doi: 10.1016/j.ymeth.2017.04.005. Epub 2017 Apr 13. Methods. 2017. PMID: 28412289 Free PMC article. Review.
-
Three-dimensional structure of the native spliceosome by cryo-electron microscopy.Mol Cell. 2004 Sep 10;15(5):833-9. doi: 10.1016/j.molcel.2004.07.022. Mol Cell. 2004. PMID: 15350226
-
Dynamic Supraspliceosomes Are Assembled on Different Transcripts Regardless of Their Intron Number and Splicing State.Front Genet. 2020 May 15;11:409. doi: 10.3389/fgene.2020.00409. eCollection 2020. Front Genet. 2020. PMID: 32499811 Free PMC article.
-
Proteomic analysis of in vivo-assembled pre-mRNA splicing complexes expands the catalog of participating factors.Nucleic Acids Res. 2007;35(12):3928-44. doi: 10.1093/nar/gkm347. Epub 2007 May 30. Nucleic Acids Res. 2007. PMID: 17537823 Free PMC article.
Cited by
-
ARiBo pull-down for riboproteomic studies based on label-free quantitative mass spectrometry.RNA. 2016 Nov;22(11):1760-1770. doi: 10.1261/rna.057513.116. Epub 2016 Sep 22. RNA. 2016. PMID: 27659051 Free PMC article.
-
A novel role of U1 snRNP: Splice site selection from a distance.Biochim Biophys Acta Gene Regul Mech. 2019 Jun;1862(6):634-642. doi: 10.1016/j.bbagrm.2019.04.004. Epub 2019 Apr 28. Biochim Biophys Acta Gene Regul Mech. 2019. PMID: 31042550 Free PMC article. Review.
-
The Chloroplast Trans-Splicing RNA-Protein Supercomplex from the Green Alga Chlamydomonas reinhardtii.Cells. 2021 Feb 1;10(2):290. doi: 10.3390/cells10020290. Cells. 2021. PMID: 33535503 Free PMC article. Review.
-
A Ribonucleoprotein Supercomplex Involved in trans-Splicing of Organelle Group II Introns.J Biol Chem. 2016 Oct 28;291(44):23330-23342. doi: 10.1074/jbc.M116.750570. Epub 2016 Sep 19. J Biol Chem. 2016. PMID: 27645995 Free PMC article.
-
The Supraspliceosome - A Multi-Task Machine for Regulated Pre-mRNA Processing in the Cell Nucleus.Comput Struct Biotechnol J. 2014 Sep 28;11(19):113-22. doi: 10.1016/j.csbj.2014.09.008. eCollection 2014 Sep. Comput Struct Biotechnol J. 2014. PMID: 25408845 Free PMC article. Review.
References
-
- Burge C.B., Tuschl T.H., Sharp P.A. In: The RNA World. 2nd ed. Gesteland R.F., Cech T.R., Atkins J.F., editors. Cold Spring Harbor Laboratory Press; New York, NY, USA: 1999. pp. 525–560.
-
- Brow D.A. Allosteric cascade of spliceosome activation. Annu. Rev. Genet. 2002;36:333–360. - PubMed
-
- Will C.L., Lührmann R. Spliceosomal UsnRNP biogenesis, structure and function. Curr. Opin. Cell Biol. 2001;13:290–301. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

