A modular view of the diversity of cell-density-encoding schemes in bacterial quorum-sensing systems
- PMID: 24988360
- PMCID: PMC4119280
- DOI: 10.1016/j.bpj.2014.05.031
A modular view of the diversity of cell-density-encoding schemes in bacterial quorum-sensing systems
Abstract
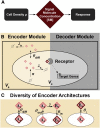
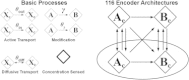
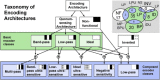
Certain environmental parameters are accessible to cells only indirectly and require an encoding step for cells to retrieve the relevant information. A prominent example is the phenomenon of quorum sensing by microorganisms, where information about cell density is encoded by means of secreted signaling molecules. The mapping of cell density to signal molecule concentration and the corresponding network modules involved have been at least partially characterized in many bacteria, and vary markedly between different systems. In this study, we investigate theoretically how differences in signal transport, signal modification, and site of signal detection shape the encoding function and affect the sensitivity and the noise characteristics of the cell-density-encoding process. We find that different modules are capable of implementing both fairly basic as well as more complex encoding schemes, whose qualitative characteristics vary with cell density and are linked to network architecture, providing the basis for a hierarchical classification scheme. We exploit the tight relationship between encoding behavior and network architecture to constrain the network topology of partially characterized natural systems, and verify one such prediction by showing experimentally that Vibrio harveyi is capable of importing Autoinducer 2. The framework developed in this research can serve not only to guide reverse engineering of natural systems but also to stimulate the design of synthetic systems and generally facilitate a better understanding of the complexities arising in the quorum-sensing process because of variations in the physical organization of the encoder network module.
Copyright © 2014 The Authors. Published by Elsevier Inc. All rights reserved.
Figures



Similar articles
-
Ultrasensitivity and noise amplification in a model of V. harveyi quorum sensing.Phys Rev E. 2016 Jun;93(6):062418. doi: 10.1103/PhysRevE.93.062418. Epub 2016 Jun 28. Phys Rev E. 2016. PMID: 27415309
-
Bacterial quorum-sensing network architectures.Annu Rev Genet. 2009;43:197-222. doi: 10.1146/annurev-genet-102108-134304. Annu Rev Genet. 2009. PMID: 19686078 Free PMC article. Review.
-
Social Evolution Selects for Redundancy in Bacterial Quorum Sensing.PLoS Biol. 2016 Feb 29;14(2):e1002386. doi: 10.1371/journal.pbio.1002386. eCollection 2016 Feb. PLoS Biol. 2016. PMID: 26927849 Free PMC article.
-
Quorum-sensing regulates biofilm formation in Vibrio scophthalmi.BMC Microbiol. 2012 Dec 3;12:287. doi: 10.1186/1471-2180-12-287. BMC Microbiol. 2012. PMID: 23198796 Free PMC article.
-
Quorum sensing in Vibrio spp.: the complexity of multiple signalling molecules in marine and aquatic environments.Crit Rev Microbiol. 2019 Aug;45(4):451-471. doi: 10.1080/1040841X.2019.1624499. Epub 2019 Jun 26. Crit Rev Microbiol. 2019. PMID: 31241379 Review.
Cited by
-
Interfering with Bacterial Quorum Sensing.Perspect Medicin Chem. 2016 Jan 18;8:1-15. doi: 10.4137/PMC.S13209. eCollection 2016. Perspect Medicin Chem. 2016. PMID: 26819549 Free PMC article.
-
The phosphorylation flow of the Vibrio harveyi quorum-sensing cascade determines levels of phenotypic heterogeneity in the population.J Bacteriol. 2015 May;197(10):1747-56. doi: 10.1128/JB.02544-14. Epub 2015 Mar 9. J Bacteriol. 2015. PMID: 25755191 Free PMC article.
-
Quadruple quorum-sensing inputs control Vibrio cholerae virulence and maintain system robustness.PLoS Pathog. 2015 Apr 15;11(4):e1004837. doi: 10.1371/journal.ppat.1004837. eCollection 2015 Apr. PLoS Pathog. 2015. PMID: 25874462 Free PMC article.
-
The quorum-sensing regulator ComA from Bacillus subtilis activates transcription using topologically distinct DNA motifs.Nucleic Acids Res. 2016 Mar 18;44(5):2160-72. doi: 10.1093/nar/gkv1242. Epub 2015 Nov 17. Nucleic Acids Res. 2016. PMID: 26582911 Free PMC article.
-
Positive Autoregulation of an Acyl-Homoserine Lactone Quorum-Sensing Circuit Synchronizes the Population Response.mBio. 2017 Jul 25;8(4):e01079-17. doi: 10.1128/mBio.01079-17. mBio. 2017. PMID: 28743819 Free PMC article.
References
-
- Fuqua C., Greenberg E.P. Listening in on bacteria: acyl-homoserine lactone signalling. Nat. Rev. Mol. Cell Biol. 2002;3:685–695. - PubMed
-
- Bassler B.L. Small talk. Cell-to-cell communication in bacteria. Cell. 2002;109:421–424. - PubMed
-
- Miller M.B., Bassler B.L. Quorum sensing in bacteria. Annu. Rev. Microbiol. 2001;55:165–199. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

