Integration of mapped RNA-Seq reads into automatic training of eukaryotic gene finding algorithm
- PMID: 24990371
- PMCID: PMC4150757
- DOI: 10.1093/nar/gku557
Integration of mapped RNA-Seq reads into automatic training of eukaryotic gene finding algorithm
Abstract
We present a new approach to automatic training of a eukaryotic ab initio gene finding algorithm. With the advent of Next-Generation Sequencing, automatic training has become paramount, allowing genome annotation pipelines to keep pace with the speed of genome sequencing. Earlier we developed GeneMark-ES, currently the only gene finding algorithm for eukaryotic genomes that performs automatic training in unsupervised ab initio mode. The new algorithm, GeneMark-ET augments GeneMark-ES with a novel method that integrates RNA-Seq read alignments into the self-training procedure. Use of 'assembled' RNA-Seq transcripts is far from trivial; significant error rate of assembly was revealed in recent assessments. We demonstrated in computational experiments that the proposed method of incorporation of 'unassembled' RNA-Seq reads improves the accuracy of gene prediction; particularly, for the 1.3 GB genome of Aedes aegypti the mean value of prediction Sensitivity and Specificity at the gene level increased over GeneMark-ES by 24.5%. In the current surge of genomic data when the need for accurate sequence annotation is higher than ever, GeneMark-ET will be a valuable addition to the narrow arsenal of automatic gene prediction tools.
© The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


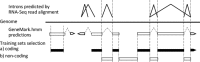

References
-
- Burge C., Karlin S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 1997;268:78–94. - PubMed
-
- Stanke M., Waack S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics. 2003;19(Suppl 2):ii215–ii225. - PubMed
-
- Majoros W.H., Pertea M., Salzberg S.L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics. 2004;20:2878–2879. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

