Specific genomic aberrations predict survival, but low mutation rate in cancer hot spots, in clear cell renal cell carcinoma
- PMID: 24992170
- PMCID: PMC4431677
- DOI: 10.1097/PAI.0000000000000087
Specific genomic aberrations predict survival, but low mutation rate in cancer hot spots, in clear cell renal cell carcinoma
Abstract
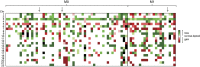
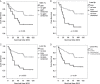
Detailed genetic profiling of clear cell renal cell carcinoma (ccRCC) has revealed genomic regions commonly affected by structural changes and a general genetic heterogeneity. VHL and PBRM1, both located at chromosome 3p, are 2 major genes mutated at high frequency but apart from these aberrations, the mutational landscape in ccRCC is largely undefined. Potential prognostic information given by the genomic changes appears to depend on the particular cohort studied. We analyzed a Swedish ccRCC cohort of 74 patients and found common changes (loss or gain occurring in >20% of the tumors) in 12 chromosomal regions (1p, 3p, 3q, 5q, 6q, 7p, 7q 8p, 9p, 9q, 10q, and 14q). A poor outcome was associated with gain of 7q and losses on 9p, 9q, and 14q. These aberrations were more frequent in metastasized tumors, suggesting alterations of genes important for tumor progression. Sequencing of 48 genes implicated in cancer revealed that only VHL, TP53, and PTEN were mutated at a noticeable frequency (51%, 9%, and 9%, respectively). Shorter relative telomere length (RTL) has been associated with loss of specific chromosomal regions in ccRCC tumors, but we could not verify this finding. However, a significantly lower tumor/nontumor (T/N) RTL ratio was detected for tumors with losses in 4q or 9p. In conclusion, poor outcome in ccRCC was associated with gain of 7q and loss on 9p, 9q, and 14q, whereas the mutation rate overall was low in a screen of cancer-associated genes.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Deletions of chromosomes 3p and 14q molecularly subclassify clear cell renal cell carcinoma.Cancer. 2013 Apr 15;119(8):1547-54. doi: 10.1002/cncr.27947. Epub 2013 Jan 18. Cancer. 2013. PMID: 23335244
-
Patterns of gene expression and copy-number alterations in von-hippel lindau disease-associated and sporadic clear cell carcinoma of the kidney.Cancer Res. 2009 Jun 1;69(11):4674-81. doi: 10.1158/0008-5472.CAN-09-0146. Cancer Res. 2009. PMID: 19470766 Free PMC article.
-
Somatic Copy Number Alterations and Associated Genes in Clear-Cell Renal-Cell Carcinoma in Brazilian Patients.Int J Mol Sci. 2021 Feb 25;22(5):2265. doi: 10.3390/ijms22052265. Int J Mol Sci. 2021. PMID: 33668731 Free PMC article.
-
A Scoping Review of Population Diversity in the Common Genomic Aberrations of Clear Cell Renal Cell Carcinoma.Oncology. 2025;103(4):341-350. doi: 10.1159/000541370. Epub 2024 Sep 9. Oncology. 2025. PMID: 39250899 Free PMC article.
-
Genomics and clinical correlates of renal cell carcinoma.World J Urol. 2018 Dec;36(12):1899-1911. doi: 10.1007/s00345-018-2429-x. Epub 2018 Aug 11. World J Urol. 2018. PMID: 30099580 Free PMC article. Review.
Cited by
-
A Review of the Paradigmatic Role of Adipose Tissue in Renal Cancer: Fat Measurement and Tumor Behavior Features.Cancers (Basel). 2024 Apr 27;16(9):1697. doi: 10.3390/cancers16091697. Cancers (Basel). 2024. PMID: 38730649 Free PMC article. Review.
-
Genomic Alteration in Head and Neck Squamous Cell Carcinoma (HNSCC) Cell Lines Inferred from Karyotyping, Molecular Cytogenetics, and Array Comparative Genomic Hybridization.PLoS One. 2016 Aug 8;11(8):e0160901. doi: 10.1371/journal.pone.0160901. eCollection 2016. PLoS One. 2016. PMID: 27501229 Free PMC article.
-
Anti-angiogenesis and Immunotherapy: Novel Paradigms to Envision Tailored Approaches in Renal Cell-Carcinoma.J Clin Med. 2020 May 24;9(5):1594. doi: 10.3390/jcm9051594. J Clin Med. 2020. PMID: 32456352 Free PMC article. Review.
-
Recent Advances in the Management of Clear Cell Renal Cell Carcinoma: Novel Biomarkers and Targeted Therapies.Cancers (Basel). 2023 Jun 16;15(12):3207. doi: 10.3390/cancers15123207. Cancers (Basel). 2023. PMID: 37370817 Free PMC article. Review.
-
DNA methylation status defines clinicopathological parameters including survival for patients with clear cell renal cell carcinoma (ccRCC).Tumour Biol. 2016 Aug;37(8):10219-28. doi: 10.1007/s13277-016-4893-5. Epub 2016 Jan 30. Tumour Biol. 2016. PMID: 26831665
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

