MicroRNA-130a is up-regulated in mouse liver by iron deficiency and targets the bone morphogenetic protein (BMP) receptor ALK2 to attenuate BMP signaling and hepcidin transcription
- PMID: 25002578
- PMCID: PMC4156078
- DOI: 10.1074/jbc.M114.577387
MicroRNA-130a is up-regulated in mouse liver by iron deficiency and targets the bone morphogenetic protein (BMP) receptor ALK2 to attenuate BMP signaling and hepcidin transcription
Abstract
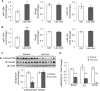
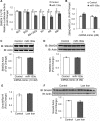
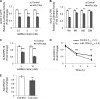
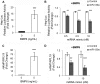
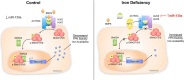
Systemic iron balance is controlled by the liver peptide hormone hepcidin, which is transcriptionally regulated by the bone morphogenetic protein (BMP)-SMAD pathway. In iron deficiency, liver BMP-SMAD signaling and hepcidin are suppressed as a compensatory mechanism to increase iron availability. MicroRNAs are small regulatory RNAs that have an increasingly recognized role in many biologic processes but are only recently implicated in iron homeostasis regulation. Here, we demonstrate that liver expression of the microRNA miR-130a is up-regulated by iron deficiency in mice. We identify the BMP6-SMAD signaling pathway as a functional target of miR-130a in hepatoma-derived Hep3B cells. Although the TGF-β/BMP common mediator SMAD4 was previously reported to be an miR-130a target to inhibit TGF-β signaling, we do not confirm SMAD4 as an miR-130a target in our biologic system. Instead, we determine that the BMP type I receptor ALK2 is a novel target of miR-130a and that miR-130a binds to two specific sites in the 3'-untranslated region to reduce ALK2 mRNA stability. Moreover, we show in mice that the increased liver miR-130a during iron deficiency is associated with reduced liver Alk2 mRNA levels. Finally, we demonstrate that down-regulation of ALK2 by miR-130a has a functional effect to inhibit BMP6-induced hepcidin transcription in Hep3B cells. Our data suggest that iron deficiency increases liver miR-130a, which, by targeting ALK2, may contribute to reduce BMP-SMAD signaling, suppress hepcidin synthesis, and thereby promote iron availability.
Keywords: ALK2; Activin A Receptor, Type I (ACVR1); Bone Morphogenetic Protein (BMP); Hepcidin; Iron Metabolism; MicroRNA (miRNA); SMAD Transcription Factor; SMAD4; Signal Transduction; miR-130a.
© 2014 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures


Similar articles
-
ALK3 undergoes ligand-independent homodimerization and BMP-induced heterodimerization with ALK2.Free Radic Biol Med. 2018 Dec;129:127-137. doi: 10.1016/j.freeradbiomed.2018.09.021. Epub 2018 Sep 15. Free Radic Biol Med. 2018. PMID: 30227271 Free PMC article.
-
Transforming Growth Factor β1 (TGF-β1) Activates Hepcidin mRNA Expression in Hepatocytes.J Biol Chem. 2016 Jun 17;291(25):13160-74. doi: 10.1074/jbc.M115.691543. Epub 2016 Apr 27. J Biol Chem. 2016. PMID: 27129231 Free PMC article.
-
A Recombinant Antibody Against ALK2 Promotes Tissue Iron Redistribution and Contributes to Anemia Resolution in a Mouse Model of Anemia of Inflammation.Am J Hematol. 2025 May;100(5):797-812. doi: 10.1002/ajh.27578. Epub 2025 Jan 10. Am J Hematol. 2025. PMID: 39791515 Free PMC article.
-
Hepcidin and the BMP-SMAD pathway: An unexpected liaison.Vitam Horm. 2019;110:71-99. doi: 10.1016/bs.vh.2019.01.004. Epub 2019 Feb 10. Vitam Horm. 2019. PMID: 30798817 Review.
-
Bone morphogenetic proteins.Growth Factors. 2004 Dec;22(4):233-41. doi: 10.1080/08977190412331279890. Growth Factors. 2004. PMID: 15621726 Review.
Cited by
-
[Progress on epigenetic regulation of iron homeostasis].Zhejiang Da Xue Xue Bao Yi Xue Ban. 2020 May 25;49(1):58-70. doi: 10.3785/j.issn.1008-9292.2020.02.25. Zhejiang Da Xue Xue Bao Yi Xue Ban. 2020. PMID: 32621410 Free PMC article. Review. Chinese.
-
Clinical implications of miRNAs in erythropoiesis, anemia, and other hematological disorders.Mol Biol Rep. 2024 Oct 18;51(1):1064. doi: 10.1007/s11033-024-09981-w. Mol Biol Rep. 2024. PMID: 39422834 Review.
-
Expression and Functional Analysis of Hepcidin from Mandarin Fish (Siniperca chuatsi).Int J Mol Sci. 2019 Nov 9;20(22):5602. doi: 10.3390/ijms20225602. Int J Mol Sci. 2019. PMID: 31717495 Free PMC article.
-
Molecular perspective of iron uptake, related diseases, and treatments.Blood Res. 2019 Mar;54(1):10-16. doi: 10.5045/br.2019.54.1.10. Epub 2019 Mar 21. Blood Res. 2019. PMID: 30956958 Free PMC article. Review.
-
HFE-Related Hemochromatosis May Be a Primary Kupffer Cell Disease.Biomedicines. 2025 Mar 10;13(3):683. doi: 10.3390/biomedicines13030683. Biomedicines. 2025. PMID: 40149659 Free PMC article. Review.
References
-
- Babitt J. L., Huang F. W., Wrighting D. M., Xia Y., Sidis Y., Samad T. A., Campagna J. A., Chung R. T., Schneyer A. L., Woolf C. J., Andrews N. C., Lin H. Y. (2006) Bone morphogenetic protein signaling by hemojuvelin regulates hepcidin expression. Nat. Genet. 38, 531–539 - PubMed
-
- Meynard D., Kautz L., Darnaud V., Canonne-Hergaux F., Coppin H., Roth M. P. (2009) Lack of the bone morphogenetic protein BMP6 induces massive iron overload. Nat. Genet. 41, 478–481 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

