The molecular basis of IL-10 function: from receptor structure to the onset of signaling
- PMID: 25004819
- PMCID: PMC4489423
- DOI: 10.1007/978-3-662-43492-5_9
The molecular basis of IL-10 function: from receptor structure to the onset of signaling
Abstract
Assembly of the cell surface IL-10 receptor complex is the first step in initiating IL-10 signaling pathways that regulate intestinal inflammation, viral persistence and even tumor surveillance. The discovery of IL-10 homologs in the genomes of herpes viruses suggests IL-10 signaling pathways can be manipulated at the level of the receptor complex. This chapter will describe our current molecular understanding of IL-10 receptor assembly based on crystal structures and biochemical analyses of cellular and viral IL-10 receptor complexes.
Figures

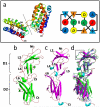


References
-
- Bazan JF. Shared architecture of hormone binidng in type I and type II interferon receptors. Cell. 1990;61:753–754. - PubMed
-
- Blackburn SD, Wherry EJ. IL-10, T cell exhaustion and viral persistence. Trends Microbiol. 2007;15:143–6. - PubMed
-
- Bleicher L, de Moura PR, Watanabe L, Colau D, Dumoutier L, Renauld JC, Polikarpov I. Crystal structure of the IL-22/IL-22R1 complex and its implications for the IL-22 signaling mechanism. FEBS Lett. 2008;582:2985–92. doi: 10.1016/j.febslet.2008.07.046. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

