The basic amino acids in the coiled-coil domain of CIN85 regulate its interaction with c-Cbl and phosphatidic acid during epidermal growth factor receptor (EGFR) endocytosis
- PMID: 25005938
- PMCID: PMC4096430
- DOI: 10.1186/1471-2091-15-13
The basic amino acids in the coiled-coil domain of CIN85 regulate its interaction with c-Cbl and phosphatidic acid during epidermal growth factor receptor (EGFR) endocytosis
Abstract
Background: During EGFR internalization CIN85 bridges EGFR-Cbl complex, endocytic machinery and fusible membrane through the interactions of CIN85 with c-Cbl, endophilins and phosphatidic acid. These protein-protein and protein-lipid interactions are mediated or regulated by the positively charged C-terminal coiled-coil domain of CIN85. However, the details of CIN85-lipid interaction remain unknown. The present study suggested a possible electric interaction between the negative charge of phosphatidic acid and the positive charge of basic amino acids in coiled-coil domain.
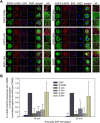
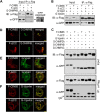
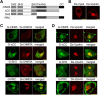
Results: Mutations of the basic amino acids in the coiled-coil domain, especially K645, K646, R648 and R650, into neutral amino acid alanine completely blocked the interaction of CIN85 with c-Cbl or phosphatidic acid. However, they did not affect CIN85-endophilin interaction. In addition, CIN85 was found to associate with the internalized EGFR endosomes. It interacted with several ESCRT (Endosomal Sorting Complex Required for Transport) component proteins for ESCRT assembly on endosomal membrane. Mutations in the coiled-coil domain (deletion of the coiled-coil domain or point mutations of the basic amino acids) dissociated CIN85 from endosomes. These mutants bound the ESCRT components in cytoplasm to prevent them from assembly on endosomal membrane and inhibited EGFR sorting for degradation.
Conclusions: As an adaptor protein, CIN85 interacts with variety of partners through several domains. The positive charges of basic amino acids in the coiled-coil domain are not only involved in the interaction with phosphatidic acid, but also regulate the interaction of CIN85 with c-Cbl. CIN85 also interacts with ESCRT components for protein sorting in endosomes. These CIN85-protein and CIN85-lipid interactions enable CIN85 to link EGFR-Cbl endocytic complex with fusible membrane during EGFR endocytosis and subsequently to facilitate ESCRT formation on endosomal membrane for EGFR sorting and degradation.
Figures




Similar articles
-
CIN85 associates with endosomal membrane and binds phosphatidic acid.Cell Res. 2009 Jun;19(6):733-46. doi: 10.1038/cr.2009.51. Cell Res. 2009. PMID: 19417776
-
SH3KBP1-binding protein 1 prevents epidermal growth factor receptor degradation by the interruption of c-Cbl-CIN85 complex.Cell Biochem Funct. 2011 Oct;29(7):589-96. doi: 10.1002/cbf.1792. Epub 2011 Aug 9. Cell Biochem Funct. 2011. PMID: 21830225 Free PMC article.
-
Sprouty2 acts at the Cbl/CIN85 interface to inhibit epidermal growth factor receptor downregulation.EMBO Rep. 2005 Jul;6(7):635-41. doi: 10.1038/sj.embor.7400453. EMBO Rep. 2005. PMID: 15962011 Free PMC article.
-
[Ubiquitination-mediated degradation of epidermal growth factor receptor].Zhongguo Yi Xue Ke Xue Yuan Xue Bao. 2005 Feb;27(1):120-7. Zhongguo Yi Xue Ke Xue Yuan Xue Bao. 2005. PMID: 15782507 Review. Chinese.
-
Endocytosis and intracellular trafficking of ErbBs.Exp Cell Res. 2008 Oct 15;314(17):3093-106. doi: 10.1016/j.yexcr.2008.08.013. Epub 2008 Aug 28. Exp Cell Res. 2008. PMID: 18793634 Free PMC article. Review.
Cited by
-
Impact of the interaction between herpes simplex virus 1 ICP22 and FACT on viral gene expression and pathogenesis.J Virol. 2024 Aug 20;98(8):e0073724. doi: 10.1128/jvi.00737-24. Epub 2024 Jul 17. J Virol. 2024. PMID: 39016551 Free PMC article.
-
New Era of Diacylglycerol Kinase, Phosphatidic Acid and Phosphatidic Acid-Binding Protein.Int J Mol Sci. 2020 Sep 16;21(18):6794. doi: 10.3390/ijms21186794. Int J Mol Sci. 2020. PMID: 32947951 Free PMC article. Review.
-
Intrinsic lipid binding activity of ATG16L1 supports efficient membrane anchoring and autophagy.EMBO J. 2019 May 2;38(9):e100554. doi: 10.15252/embj.2018100554. Epub 2019 Apr 1. EMBO J. 2019. PMID: 30936093 Free PMC article.
-
ARAP3 functions in hematopoietic stem cells.PLoS One. 2014 Dec 26;9(12):e116107. doi: 10.1371/journal.pone.0116107. eCollection 2014. PLoS One. 2014. PMID: 25542002 Free PMC article.
-
EGFR mutations and abnormal trafficking in cancers.Mol Biol Rep. 2024 Aug 21;51(1):924. doi: 10.1007/s11033-024-09865-z. Mol Biol Rep. 2024. PMID: 39167290 Review.
References
-
- Dikic I. CIN85/CMS family of adaptor molecules. FEBS Lett. 2002;529:110–115. - PubMed
-
- Take H, Watanabe S, Takeda K, Yu ZX, Iwata N, Kajigaya S. Cloning and characterization of a novel adaptor protein, CIN85, that interacts with c-Cbl. Biochem Biophys Res Commun. 2000;268:321–328. - PubMed
-
- Soubeyran P, Kowanetz K, Szymkiewicz I, Langdon WY, Dikic I. Cbl-CIN85-endophilin complex mediates ligand-induced downregulation of EGF receptors. Nature. 2002;416:183–187. - PubMed
-
- Petrelli A, Gilestro GF, Lanzardo S, Comoglio PM, Migone N, Giordano S. The endophilin-CIN85-Cbl complex mediates ligand-dependent downregulation of c-Met. Nature. 2002;416:187–190. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

