Loss of MITF expression during human embryonic stem cell differentiation disrupts retinal pigment epithelium development and optic vesicle cell proliferation
- PMID: 25008112
- PMCID: PMC4222367
- DOI: 10.1093/hmg/ddu351
Loss of MITF expression during human embryonic stem cell differentiation disrupts retinal pigment epithelium development and optic vesicle cell proliferation
Abstract
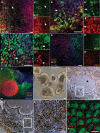
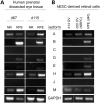
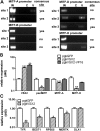
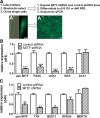
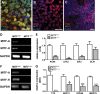
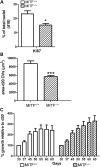
Microphthalmia-associated transcription factor (MITF) is a master regulator of pigmented cell survival and differentiation with direct transcriptional links to cell cycle, apoptosis and pigmentation. In mouse, Mitf is expressed early and uniformly in optic vesicle (OV) cells as they evaginate from the developing neural tube, and null Mitf mutations result in microphthalmia and pigmentation defects. However, homozygous mutations in MITF have not been identified in humans; therefore, little is known about its role in human retinogenesis. We used a human embryonic stem cell (hESC) model that recapitulates numerous aspects of retinal development, including OV specification and formation of retinal pigment epithelium (RPE) and neural retina progenitor cells (NRPCs), to investigate the earliest roles of MITF. During hESC differentiation toward a retinal lineage, a subset of MITF isoforms was expressed in a sequence and tissue distribution similar to that observed in mice. In addition, we found that promoters for the MITF-A, -D and -H isoforms were directly targeted by Visual Systems Homeobox 2 (VSX2), a transcription factor involved in patterning the OV toward a NRPC fate. We then manipulated MITF RNA and protein levels at early developmental stages and observed decreased expression of eye field transcription factors, reduced early OV cell proliferation and disrupted RPE maturation. This work provides a foundation for investigating MITF and other highly complex, multi-purposed transcription factors in a dynamic human developmental model system.
© The Author 2014. Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oup.com.
Figures

References
-
- Zuber M., Gestri G., Viczian A., Barsacchi G., Harris W. Specification of the vertebrate eye by a network of eye field transcription factors. Development. 2003;130:5155–5167. - PubMed
-
- Fuhrman S., Zou C., Levine E. Retinal pigment epithelium development, plasticity and tissue homeostasis. Exp. Eye Res. 2013 doi:10.1016/j.exer.2013.09.003. - DOI - PMC - PubMed
-
- Nguyen M., Arnheiter H. Signaling and transcriptional regulation in early mammalian eye development: a link between FGF and MITF. Development. 2000;127:3581–3591. - PubMed
-
- Bora N., Conway S., Liang H., Smith S. Transient overexpression of the Microphthalmia gene in the eyes of Microphthalmia vitiligo mutant mice. Dev. Dyn. 1998;213:283–292. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

