Autographa californica multiple nucleopolyhedrovirus DNA polymerase C terminus is required for nuclear localization and viral DNA replication
- PMID: 25008932
- PMCID: PMC4178887
- DOI: 10.1128/JVI.01167-14
Autographa californica multiple nucleopolyhedrovirus DNA polymerase C terminus is required for nuclear localization and viral DNA replication
Abstract
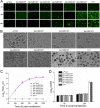
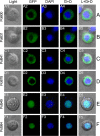
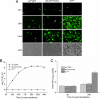
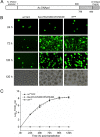
The DNA polymerase (DNApol) of the baculovirus Autographa californica multiple nucleopolyhedrovirus (AcMNPV) is essential for viral DNA replication. The DNApol exonuclease and polymerase domains are highly conserved and are considered functional in DNA replication. However, the role of the DNApol C terminus has not yet been characterized. To identify whether only the exonuclease and polymerase domains are sufficient for viral DNA replication, several DNApol C-terminal truncations were cloned into a dnapol-null AcMNPV bacmid with a green fluorescent protein (GFP) reporter. Surprisingly, most of the truncation constructs, despite containing both exonuclease and polymerase domains, could not rescue viral DNA replication and viral production in bacmid-transfected Sf21 cells. Moreover, GFP fusions of these same truncations failed to localize to the nucleus. Truncation of the C-terminal amino acids 950 to 984 showed nuclear localization but allowed for only limited and delayed viral spread. The C terminus contains a typical bipartite nuclear localization signal (NLS) motif at residues 804 to 827 and a monopartite NLS motif at residues 939 to 948. Each NLS, as a GFP fusion peptide, localized to the nucleus, but both NLSs were required for nuclear localization of DNApol. Alanine substitutions in a highly conserved baculovirus DNApol sequence at AcMNPV DNApol amino acids 972 to 981 demonstrated its importance for virus production and DNA replication. Collectively, the data indicated that the C terminus of AcMNPV DNApol contains two NLSs and a conserved motif, all of which are required for nuclear localization of DNApol, viral DNA synthesis, and virus production.
Importance: The baculovirus DNA polymerase (DNApol) is a highly specific polymerase that allows viral DNA synthesis and hence virus replication in infected insect cells. We demonstrated that the exonuclease and polymerase domains of Autographa californica multiple nucleopolyhedrovirus (AcMNPV) alone are insufficient for viral DNA synthesis and virus replication. Rather, we identified three features, including two nuclear localization signals and a highly conserved 10-amino-acid sequence in the AcMNPV DNApol C terminus, all three of which are important for both nuclear localization of DNApol and for DNApol activity, as measured by viral DNA synthesis and virus replication.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures



Similar articles
-
The deficiency in nuclear localization signal of Neodiprion lecontei nucleopolyhedrovirus DNA polymerase prevents rescue of viral DNA replication and virus production in dnapol-null Autographa californica multiple nucleopolyhedrovirus.Virus Res. 2019 Jun;266:52-57. doi: 10.1016/j.virusres.2019.04.005. Epub 2019 Apr 13. Virus Res. 2019. PMID: 30991090
-
The N Terminus of Autographa californica Multiple Nucleopolyhedrovirus DNA Polymerase Is Required for Efficient Viral DNA Replication and Virus and Occlusion Body Production.J Virol. 2018 May 14;92(11):e00398-18. doi: 10.1128/JVI.00398-18. Print 2018 Jun 1. J Virol. 2018. PMID: 29563284 Free PMC article.
-
Identification and Characterization of the Nucleolar Localization Signal of Autographa californica Multiple Nucleopolyhedrovirus LEF5.J Virol. 2020 Jan 31;94(4):e01891-19. doi: 10.1128/JVI.01891-19. Print 2020 Jan 31. J Virol. 2020. PMID: 31776271 Free PMC article.
-
me53 encoded by Autographa californica multiple nucleopolyhedrovirus: from mechanism to function.Virus Genes. 2023 Apr;59(2):188-194. doi: 10.1007/s11262-022-01943-3. Epub 2022 Oct 13. Virus Genes. 2023. PMID: 36229721 Review.
-
Baculoviruses as gene therapy vectors for human prostate cancer.J Invertebr Pathol. 2011 Jul;107 Suppl:S59-70. doi: 10.1016/j.jip.2011.05.006. J Invertebr Pathol. 2011. PMID: 21784232 Review.
Cited by
-
Nuclear Translocation Sequence and Region in Autographa californica Multiple Nucleopolyhedrovirus ME53 That Are Important for Optimal Baculovirus Production.J Virol. 2016 Mar 28;90(8):3953-3965. doi: 10.1128/JVI.03115-15. Print 2016 Apr. J Virol. 2016. PMID: 26842471 Free PMC article.
-
Autographa Californica Multiple Nucleopolyhedrovirus orf13 Is Required for Efficient Nuclear Egress of Nucleocapsids.Virol Sin. 2021 Oct;36(5):968-980. doi: 10.1007/s12250-021-00353-3. Epub 2021 Mar 15. Virol Sin. 2021. PMID: 33721216 Free PMC article.
-
The Motif of 76KRKCSK in Bm65 Is an Efficient Nuclear Localization Signal Involved in Production of Infectious Virions.Front Microbiol. 2019 Nov 26;10:2739. doi: 10.3389/fmicb.2019.02739. eCollection 2019. Front Microbiol. 2019. PMID: 32038506 Free PMC article.
-
Improving Baculovirus Infectivity by Efficiently Embedding Enhancing Factors into Occlusion Bodies.Appl Environ Microbiol. 2017 Jun 30;83(14):e00595-17. doi: 10.1128/AEM.00595-17. Print 2017 Jul 15. Appl Environ Microbiol. 2017. PMID: 28500037 Free PMC article.
-
Mechanisms Mediating Nuclear Trafficking Involved in Viral Propagation by DNA Viruses.Viruses. 2019 Nov 7;11(11):1035. doi: 10.3390/v11111035. Viruses. 2019. PMID: 31703327 Free PMC article. Review.
References
-
- King AM, Elliot E, Adam M, Carstens EB. 2012. Virus taxonomy: ninth report of the International Committee on Taxonomy of Viruses. Academic Press, New York, NY
-
- Federici BA. 1997. Baculovirus pathogenesis, p 33–56 In Miller LK. (ed), The baculoviruses. Plenum Press, New York, NY
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

