Capturing intra-tumor genetic heterogeneity by de novo mutation profiling of circulating cell-free tumor DNA: a proof-of-principle
- PMID: 25009010
- PMCID: PMC6276937
- DOI: 10.1093/annonc/mdu239
Capturing intra-tumor genetic heterogeneity by de novo mutation profiling of circulating cell-free tumor DNA: a proof-of-principle
Erratum in
-
Capturing intra-tumor genetic heterogeneity by de novo mutation profiling of circulating cell-free tumor DNA: a proof-of-principle.Ann Oncol. 2018 Nov 1;29(11):2268. doi: 10.1093/annonc/mdx804. Ann Oncol. 2018. PMID: 29718117 Free PMC article. No abstract available.
Abstract
Background: Plasma-derived cell-free tumor DNA (ctDNA) constitutes a potential surrogate for tumor DNA obtained from tissue biopsies. We posit that massively parallel sequencing (MPS) analysis of ctDNA may help define the repertoire of mutations in breast cancer and monitor tumor somatic alterations during the course of targeted therapy.
Patient and methods: A 66-year-old patient presented with synchronous estrogen receptor-positive/HER2-negative, highly proliferative, grade 2, mixed invasive ductal-lobular carcinoma with bone and liver metastases at diagnosis. DNA extracted from archival tumor material, plasma and peripheral blood leukocytes was subjected to targeted MPS using a platform comprising 300 cancer genes known to harbor actionable mutations. Multiple plasma samples were collected during the fourth line of treatment with an AKT inhibitor.
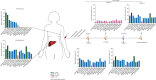
Results: Average read depths of 287x were obtained from the archival primary tumor, 139x from the liver metastasis and between 200x and 900x from ctDNA samples. Sixteen somatic non-synonymous mutations were detected in the liver metastasis, of which 9 (CDKN2A, AKT1, TP53, JAK3, TSC1, NF1, CDH1, MML3 and CTNNB1) were also detected in >5% of the alleles found in the primary tumor sample. Not all mutations identified in the metastasis were reliably identified in the primary tumor (e.g. FLT4). Analysis of ctDNA, nevertheless, captured all mutations present in the primary tumor and/or liver metastasis. In the longitudinal monitoring of the patient, the mutant allele fractions identified in ctDNA samples varied over time and mirrored the pharmacodynamic response to the targeted therapy as assessed by positron emission tomography-computed tomography.
Conclusions: This proof-of-principle study is one of the first to demonstrate that high-depth targeted MPS of plasma-derived ctDNA constitutes a potential tool for de novo mutation identification and monitoring of somatic genetic alterations during the course of targeted therapy, and may be employed to overcome the challenges posed by intra-tumor genetic heterogeneity.
Registered clinical trial: www.clinicaltrials.gov, NCT01090960.
Keywords: breast cancer; cell-free tumor DNA; disease monitoring; intra-tumor heterogeneity; massively parallel sequencing.
© The Author 2014. Published by Oxford University Press on behalf of the European Society for Medical Oncology. All rights reserved. For permissions, please email: journals.permissions@oup.com.
Figures

References
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

