Transcription factor regulation and chromosome dynamics during pseudohyphal growth
- PMID: 25009286
- PMCID: PMC4148256
- DOI: 10.1091/mbc.E14-04-0871
Transcription factor regulation and chromosome dynamics during pseudohyphal growth
Abstract
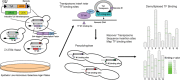
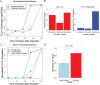
Pseudohyphal growth is a developmental pathway seen in some strains of yeast in which cells form multicellular filaments in response to environmental stresses. We used multiplexed transposon "Calling Cards" to record the genome-wide binding patterns of 28 transcription factors (TFs) in nitrogen-starved yeast. We identified TF targets relevant for pseudohyphal growth, producing a detailed map of its regulatory network. Using tools from graph theory, we identified 14 TFs that lie at the center of this network, including Flo8, Mss11, and Mfg1, which bind as a complex. Surprisingly, the DNA-binding preferences for these key TFs were unknown. Using Calling Card data, we predicted the in vivo DNA-binding motif for the Flo8-Mss11-Mfg1 complex and validated it using a reporter assay. We found that this complex binds several important targets, including FLO11, at both their promoter and termination sequences. We demonstrated that this binding pattern is the result of DNA looping, which regulates the transcription of these targets and is stabilized by an interaction with the nuclear pore complex. This looping provides yeast cells with a transcriptional memory, enabling them more rapidly to execute the filamentous growth program when nitrogen starved if they had been previously exposed to this condition.
© 2014 Mayhew and Mitra. This article is distributed by The American Society for Cell Biology under license from the author(s). Two months after publication it is available to the public under an Attribution–Noncommercial–Share Alike 3.0 Unported Creative Commons License (http://creativecommons.org/licenses/by-nc-sa/3.0).
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

