Prospects for inhibiting the post-transcriptional regulation of gene expression in hepatitis B virus
- PMID: 25009369
- PMCID: PMC4081668
- DOI: 10.3748/wjg.v20.i25.7993
Prospects for inhibiting the post-transcriptional regulation of gene expression in hepatitis B virus
Abstract
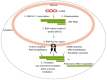
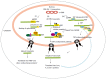
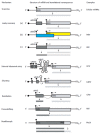
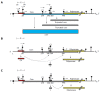
There is a continuing need for novel antivirals to treat hepatitis B virus (HBV) infection, as it remains a major health problem worldwide. Ideally new classes of antivirals would target multiple steps in the viral lifecycle. In this review, we consider the steps in which HBV RNAs are processed, exported from the nucleus and translated. These are often overlooked steps in the HBV life-cycle. HBV, like retroviruses, incorporates a number of unusual steps in these processes, which use a combination of viral and host cellular machinery. Some of these unusual steps deserve a closer scrutiny. They may provide alternative targets to existing antiviral therapies, which are associated with increasing drug resistance. The RNA post-transcriptional regulatory element identified 20 years ago promotes nucleocytoplasmic export of all unspliced HBV RNAs. There is evidence that inhibition of this step is part of the antiviral action of interferon. Similarly, the structured RNA epsilon element situated at the 5' end of the polycistronic HBV pregenomic RNA also performs key roles during HBV replication. The pregenomic RNA, which is the template for translation of both the viral core and polymerase proteins, is also encapsidated and used in replication. This complex process, regulated at the epsilon element, also presents an attractive antiviral target. These RNA elements that mediate and regulate gene expression are highly conserved and could be targeted using novel strategies employing RNAi, miRNAs or aptamers. Such approaches targeting these functionally constrained genomic regions should avoid escape mutations. Therefore understanding these regulatory elements, along with providing potential targets, may also facilitate the development of other new classes of antiviral drugs.
Keywords: Antiviral; Hepatitis B virus; Nuclear export; Nucleocytoplasmic export; Post-transcriptional control; Translational control.
Figures

Similar articles
-
A conserved RNA structural element within the hepatitis B virus post-transcriptional regulatory element enhance nuclear export of intronless transcripts and repress the splicing mechanism.Mol Biol Rep. 2015 Dec;42(12):1603-14. doi: 10.1007/s11033-015-3928-0. Epub 2015 Oct 29. Mol Biol Rep. 2015. PMID: 26514143
-
[Post-transcriptional regulation mechanism and antiviral strategy of hepatitis B virus RNA].Zhonghua Gan Zang Bing Za Zhi. 2024 May 20;32(5):474-480. doi: 10.3760/cma.j.cn501113-20240410-00191. Zhonghua Gan Zang Bing Za Zhi. 2024. PMID: 38858198 Review. Chinese.
-
Identification and characterization of a novel hepatitis B virus pregenomic RNA encapsidation inhibitor.Antiviral Res. 2020 Mar;175:104709. doi: 10.1016/j.antiviral.2020.104709. Epub 2020 Jan 12. Antiviral Res. 2020. PMID: 31940474
-
Antiviral activity and hepatoprotection by heme oxygenase-1 in hepatitis B virus infection.Gastroenterology. 2007 Oct;133(4):1156-65. doi: 10.1053/j.gastro.2007.07.021. Epub 2007 Jul 25. Gastroenterology. 2007. PMID: 17919491
-
Revisiting Hepatitis B Virus: Challenges of Curative Therapies.J Virol. 2019 Sep 30;93(20):e01032-19. doi: 10.1128/JVI.01032-19. Print 2019 Oct 15. J Virol. 2019. PMID: 31375584 Free PMC article. Review.
Cited by
-
The Dihydroquinolizinone Compound RG7834 Inhibits the Polyadenylase Function of PAPD5 and PAPD7 and Accelerates the Degradation of Matured Hepatitis B Virus Surface Protein mRNA.Antimicrob Agents Chemother. 2020 Dec 16;65(1):e00640-20. doi: 10.1128/AAC.00640-20. Print 2020 Dec 16. Antimicrob Agents Chemother. 2020. PMID: 33046485 Free PMC article.
-
A conserved RNA structural element within the hepatitis B virus post-transcriptional regulatory element enhance nuclear export of intronless transcripts and repress the splicing mechanism.Mol Biol Rep. 2015 Dec;42(12):1603-14. doi: 10.1007/s11033-015-3928-0. Epub 2015 Oct 29. Mol Biol Rep. 2015. PMID: 26514143
-
Phosphorylation and Alternative Translation on Wheat Germ Cell-Free Protein Synthesis of the DHBV Large Envelope Protein.Front Mol Biosci. 2019 Dec 3;6:138. doi: 10.3389/fmolb.2019.00138. eCollection 2019. Front Mol Biosci. 2019. PMID: 31850370 Free PMC article.
-
Osteopetrosis-Associated Transmembrane Protein 1 Recruits RNA Exosome To Restrict Hepatitis B Virus Replication.J Virol. 2020 May 18;94(11):e01800-19. doi: 10.1128/JVI.01800-19. Print 2020 May 18. J Virol. 2020. PMID: 32188736 Free PMC article.
-
Know Your Enemy: Successful Bioinformatic Approaches to Predict Functional RNA Structures in Viral RNAs.Front Microbiol. 2018 Jan 4;8:2582. doi: 10.3389/fmicb.2017.02582. eCollection 2017. Front Microbiol. 2018. PMID: 29354101 Free PMC article. Review.
References
-
- Ott JJ, Stevens GA, Groeger J, Wiersma ST. Global epidemiology of hepatitis B virus infection: new estimates of age-specific HBsAg seroprevalence and endemicity. Vaccine. 2012;30:2212–2219. - PubMed
-
- Tujios SR, Lee WM. Update in the management of chronic hepatitis B. Curr Opin Gastroenterol. 2013;29:250–256. - PubMed
-
- Zoulim F, Locarnini S. Management of treatment failure in chronic hepatitis B. J Hepatol. 2012;56 Suppl 1:S112–S122. - PubMed
-
- Lucifora J, Zoulim F. The life cycle of hepatitis B virus and antiviral targets. Future Virology. 2011;6:599–614.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

