Subtractive hybridization-mediated analysis of genes and in silico prediction of associated microRNAs under waterlogged conditions in sugarcane (Saccharum spp.)
- PMID: 25009768
- PMCID: PMC4087145
- DOI: 10.1016/j.fob.2014.05.007
Subtractive hybridization-mediated analysis of genes and in silico prediction of associated microRNAs under waterlogged conditions in sugarcane (Saccharum spp.)
Abstract
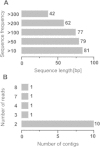
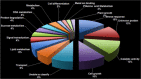
Sugarcane is an important tropical cash crop meeting 75% of world sugar demand and it is fast becoming an energy crop for the production of bio-fuel ethanol. A considerable area under sugarcane is prone to waterlogging which adversely affects both cane productivity and quality. In an effort to elucidate the genes underlying plant responses to waterlogging, a subtractive cDNA library was prepared from leaf tissue. cDNA clones were sequenced and annotated for their putative functions. Major groups of ESTs were related to stress (15%), catalytic activity (13%), cell growth (10%) and transport related proteins (6%). A few stress-related genes were identified, including senescence-associated protein, dehydration-responsive family protein, and heat shock cognate 70 kDa protein. A bioinformatics search was carried out to discover novel microRNAs (miRNAs) that can be regulated in sugarcane plants subjected to waterlogging stress. Taking advantage of the presence of miRNA precursors in the related sorghum genome, seven candidate mature miRNAs were identified in sugarcane. The application of subtraction technology allowed the identification of differentially expressed sequences and novel miRNAs in sugarcane under waterlogging stress. The comparative global transcript profiling in sugarcane plants undertaken in this study suggests that proteins associated with stress response, signal transduction, metabolic activity and ion transport play important role in conferring waterlogging tolerance in sugarcane.
Keywords: ABA, abscisic acid; ADF, actin depolymerizing factor; EST; ESTs, expressed sequence tags; Hsps, heat shock proteins; Novel miRNAs; PTGS, post-transcriptional gene silencing; REMs, remorins; SSA, snow pea pod senescence-associated; SUCEST, sugarcane EST database; Sugarcane; Suppression subtractive hybridization; TGS, transcriptional gene silencing; Waterlogging stress.
Figures

Similar articles
-
microRNAs associated with drought response in the bioenergy crop sugarcane (Saccharum spp.).PLoS One. 2012;7(10):e46703. doi: 10.1371/journal.pone.0046703. Epub 2012 Oct 11. PLoS One. 2012. PMID: 23071617 Free PMC article.
-
Expression analysis of sugarcane shaggy-like kinase (SuSK) gene identified through cDNA subtractive hybridization in sugarcane (Saccharum officinarum L.).Protoplasma. 2011 Jul;248(3):613-21. doi: 10.1007/s00709-010-0207-8. Epub 2010 Sep 18. Protoplasma. 2011. PMID: 20853012
-
Identification of genes responsive to the application of ethanol on sugarcane leaves.Plant Cell Rep. 2007 Dec;26(12):2119-28. doi: 10.1007/s00299-007-0430-8. Epub 2007 Aug 16. Plant Cell Rep. 2007. PMID: 17701412
-
MicroRNAs and Their Regulatory Role in Sugarcane.Front Plant Sci. 2017 Jun 13;8:997. doi: 10.3389/fpls.2017.00997. eCollection 2017. Front Plant Sci. 2017. PMID: 28659947 Free PMC article. Review.
-
Current breeding and genomic approaches to enhance the cane and sugar productivity under abiotic stress conditions.3 Biotech. 2020 Oct;10(10):440. doi: 10.1007/s13205-020-02416-w. Epub 2020 Sep 18. 3 Biotech. 2020. PMID: 33014683 Free PMC article. Review.
Cited by
-
Suppression Subtractive Hybridization Versus Next-Generation Sequencing in Plant Genetic Engineering: Challenges and Perspectives.Mol Biotechnol. 2015 Oct;57(10):880-903. doi: 10.1007/s12033-015-9884-z. Mol Biotechnol. 2015. PMID: 26271955 Review.
-
Differentially Expressed microRNAs and Target Genes Associated with Plastic Internode Elongation in Alternanthera philoxeroides in Contrasting Hydrological Habitats.Front Plant Sci. 2017 Dec 5;8:2078. doi: 10.3389/fpls.2017.02078. eCollection 2017. Front Plant Sci. 2017. PMID: 29259617 Free PMC article.
-
Characterization of leaf transcriptome, development and utilization of unigenes-derived microsatellite markers in sugarcane (Saccharum sp. hybrid).Physiol Mol Biol Plants. 2018 Jul;24(4):665-682. doi: 10.1007/s12298-018-0563-y. Epub 2018 Jun 4. Physiol Mol Biol Plants. 2018. PMID: 30042621 Free PMC article.
-
iTRAQ Proteomic Analysis of Wheat (Triticum aestivum L.) Genotypes Differing in Waterlogging Tolerance.Front Plant Sci. 2022 Apr 25;13:890083. doi: 10.3389/fpls.2022.890083. eCollection 2022. Front Plant Sci. 2022. PMID: 35548301 Free PMC article.
-
Transcriptional profiling in pearl millet (Pennisetum glaucum L.R. Br.) for identification of differentially expressed drought responsive genes.Physiol Mol Biol Plants. 2015 Apr;21(2):187-96. doi: 10.1007/s12298-015-0287-1. Epub 2015 Apr 15. Physiol Mol Biol Plants. 2015. PMID: 25964713 Free PMC article.
References
-
- Grivet L., Arruda P. Sugarcane genomics: depicting the complex genome of an important tropical crop. Curr. Opin. Plant Biol. 2002;5:122–127. - PubMed
-
- Extension publication, (2010) Effect of Waterlogging in Sugarcane and its Management, Sugarcane Breeding Institute India. 185.
-
- Gilbert R.A., Rainbolt C.R., Morris D.R., Bennett A.C. Morphological responses of sugarcane to long-term flooding. Agron. J. 2007;99:1622–1628.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

