Structure, stability and behaviour of nucleic acids in ionic liquids
- PMID: 25013178
- PMCID: PMC4132699
- DOI: 10.1093/nar/gku499
Structure, stability and behaviour of nucleic acids in ionic liquids
Abstract
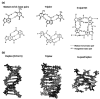
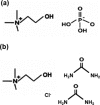
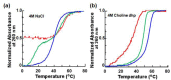
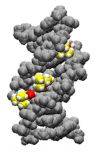
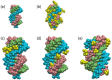
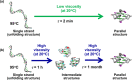
Nucleic acids have become a powerful tool in nanotechnology because of their conformational polymorphism. However, lack of a medium in which nucleic acid structures exhibit long-term stability has been a bottleneck. Ionic liquids (ILs) are potential solvents in the nanotechnology field. Hydrated ILs, such as choline dihydrogen phosphate (choline dhp) and deep eutectic solvent (DES) prepared from choline chloride and urea, are 'green' solvents that ensure long-term stability of biomolecules. An understanding of the behaviour of nucleic acids in hydrated ILs is necessary for developing DNA materials. We here review current knowledge about the structures and stabilities of nucleic acids in choline dhp and DES. Interestingly, in choline dhp, A-T base pairs are more stable than G-C base pairs, the reverse of the situation in buffered NaCl solution. Moreover, DNA triplex formation is markedly stabilized in hydrated ILs compared with aqueous solution. In choline dhp, the stability of Hoogsteen base pairs is comparable to that of Watson-Crick base pairs. Moreover, the parallel form of the G-quadruplex is stabilized in DES compared with aqueous solution. The behaviours of various DNA molecules in ILs detailed here should be useful for designing oligonucleotides for the development of nanomaterials and nanodevices.
© The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



Similar articles
-
Comparable stability of Hoogsteen and Watson-Crick base pairs in ionic liquid choline dihydrogen phosphate.Sci Rep. 2014 Jan 8;4:3593. doi: 10.1038/srep03593. Sci Rep. 2014. PMID: 24399194 Free PMC article.
-
Biological and nanotechnological applications using interactions between ionic liquids and nucleic acids.Biophys Rev. 2018 Jun;10(3):931-940. doi: 10.1007/s12551-018-0422-7. Epub 2018 Apr 23. Biophys Rev. 2018. PMID: 29687271 Free PMC article. Review.
-
Recent progress in G-quadruplex DNA in deep eutectic solvent.Methods. 2013 Nov;64(1):52-8. doi: 10.1016/j.ymeth.2013.04.017. Epub 2013 Apr 27. Methods. 2013. PMID: 23628945 Review.
-
DNA sensor's selectivity enhancement and protection from contaminating nucleases due to a hydrated ionic liquid.Analyst. 2015 Jul 7;140(13):4393-8. doi: 10.1039/c5an00545k. Analyst. 2015. PMID: 25919083
-
DNA nanotechnology in ionic liquids and deep eutectic solvents.Crit Rev Biotechnol. 2024 Aug;44(5):941-961. doi: 10.1080/07388551.2023.2229950. Epub 2023 Jul 30. Crit Rev Biotechnol. 2024. PMID: 37518062 Review.
Cited by
-
Ionic liquids: prospects for nucleic acid handling and delivery.Nucleic Acids Res. 2021 Feb 22;49(3):1201-1234. doi: 10.1093/nar/gkaa1280. Nucleic Acids Res. 2021. PMID: 33476366 Free PMC article. Review.
-
Utilising a Clinical Metabolomics LC-MS Study to Determine the Integrity of Biological Samples for Statistical Modelling after Long Term -80 °C Storage: A TOFI_Asia Sub-Study.Metabolites. 2024 May 29;14(6):313. doi: 10.3390/metabo14060313. Metabolites. 2024. PMID: 38921448 Free PMC article.
-
Ionic-Liquid-Mediated Extraction and Separation Processes for Bioactive Compounds: Past, Present, and Future Trends.Chem Rev. 2017 May 24;117(10):6984-7052. doi: 10.1021/acs.chemrev.6b00550. Epub 2017 Feb 2. Chem Rev. 2017. PMID: 28151648 Free PMC article. Review.
-
Whole genome experimental maps of DNA G-quadruplexes in multiple species.Nucleic Acids Res. 2019 May 7;47(8):3862-3874. doi: 10.1093/nar/gkz179. Nucleic Acids Res. 2019. PMID: 30892612 Free PMC article.
-
Choline Dihydrogen Phosphate Destabilizes G-Quadruplexes and Enhances Transcription Efficiency In Vitro and in Cells.ACS Omega. 2024 Jan 23;9(5):5675-5682. doi: 10.1021/acsomega.3c08087. eCollection 2024 Feb 6. ACS Omega. 2024. PMID: 38343917 Free PMC article.
References
-
- Heller M.J. DNA microarray technology: devices, systems, and applications. Annu. Rev. Biomed. Eng. 2002;4:129–153. - PubMed
-
- Ott J., Hoh J. Set association analysis of SNP case-control and microarray data. J. Comput. Biol. 2003;10:569–574. - PubMed
-
- DeSantis T.Z., Dubosarskiy I., Murray S.R., Andersen G.L. Comprehensive aligned sequence construction for automated design of effective probes (CASCADE-P) using 16S rDNA. Bioinformatics. 2003;19:1461–1468. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

