Engineering light-inducible nuclear localization signals for precise spatiotemporal control of protein dynamics in living cells
- PMID: 25019686
- PMCID: PMC4104460
- DOI: 10.1038/ncomms5404
Engineering light-inducible nuclear localization signals for precise spatiotemporal control of protein dynamics in living cells
Abstract
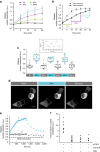
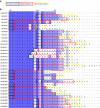
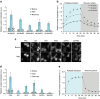
The function of many eukaryotic proteins is regulated by highly dynamic changes in their nucleocytoplasmic distribution. The ability to precisely and reversibly control nuclear translocation would, therefore, allow dissecting and engineering cellular networks. Here we develop a genetically encoded, light-inducible nuclear localization signal (LINuS) based on the LOV2 domain of Avena sativa phototropin 1. LINuS is a small, versatile tag, customizable for different proteins and cell types. LINuS-mediated nuclear import is fast and reversible, and can be tuned at different levels, for instance, by introducing mutations that alter AsLOV2 domain photo-caging properties or by selecting nuclear localization signals (NLSs) of various strengths. We demonstrate the utility of LINuS in mammalian cells by controlling gene expression and entry into mitosis with blue light.
Figures




References
-
- Werner S. L., Barken D. & Hoffmann A. Stimulus specificity of gene expression programs determined by temporal control of IKK activity. Science 309, 1857–1861 (2005). - PubMed
-
- Nelson D. E. et al. Oscillations in NF-kappaB signaling control the dynamics of gene expression. Science 306, 704–708 (2004). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

