Seed sequence-matched controls reveal limitations of small interfering RNA knockdown in functional and structural studies of hepatitis C virus NS5A-MOBKL1B interaction
- PMID: 25031347
- PMCID: PMC4178819
- DOI: 10.1128/JVI.01582-14
Seed sequence-matched controls reveal limitations of small interfering RNA knockdown in functional and structural studies of hepatitis C virus NS5A-MOBKL1B interaction
Abstract
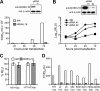
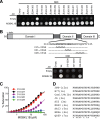
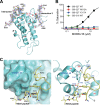
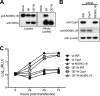
Hepatitis C virus (HCV) is a widespread human pathogen causing liver cirrhosis and cancer. Similar to the case for other viruses, HCV depends on host and viral factors to complete its life cycle. We used proteomic and yeast two-hybrid approaches to elucidate host factors involved in HCV nonstructural protein NS5A function and found that MOBKL1B interacts with NS5A. Initial experiments with small interfering RNA (siRNA) knockdown suggesting a role in HCV replication led us to examine the interaction using biochemical and structural approaches. As revealed by a cocrystal structure of a core MOBKL1B-NS5A peptide complex at 1.95 Å, NS5A binds to a hydrophobic patch on the MOBKL1B surface. Biosensor binding assays identified a highly conserved, 18-amino-acid binding site in domain II of NS5A, which encompasses residues implicated in cyclophilin A (CypA)-dependent HCV RNA replication. However, a CypA-independent HCV variant had reduced replication in MOBKL1B knockdown cells, even though its NS5A does not interact with MOBKL1B. These discordant results prompted more extensive studies of MOBKL1B gene knockdowns, which included additional siRNAs and specifically matched seed sequence siRNA controls. We found that reduced virus replication after treating cells with MOBKL1B siRNA was actually due to off-target inhibition, which indicated that the initial finding of virus replication dependence on the MOBKL1B-NS5A interaction was incorrect. Ultimately, using several approaches, we found no relationship of the MOBKL1B-NS5A interaction to virus replication. These findings collectively serve as a reminder to investigators and scientific reviewers of the pervasive impact of siRNA off-target effects on interpretation of biological data.
Importance: Our study illustrates an underappreciated shortcoming of siRNA gene knockdown technology. We initially identified a cellular protein, MOBKL1B, as a binding partner with the NS5A protein of hepatitis C virus (HCV). MOBKL1B siRNA, but not irrelevant RNA, treatment was associated with both reduced virus replication and the absence of MOBKL1B. Believing that HCV replication depended on the MOBKL1B-NS5A interaction, we carried out structural and biochemical analyses. Unexpectedly, an HCV variant lacking the MOBKL1B-NS5A interaction could not replicate after cells were treated with MOBKL1B siRNA. By repeating the MOBKL1B siRNA knockdowns and including seed sequence-matched siRNA instead of irrelevant siRNA as a control, we found that the MOBKL1B siRNAs utilized had off-target inhibitory effects on virus replication. Collectively, our results suggest that stricter controls must be utilized in all RNA interference (RNAi)-mediated gene knockdown experiments to ensure sound conclusions and a reliable scientific knowledge database.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures

Similar articles
-
Vinexin β Interacts with Hepatitis C Virus NS5A, Modulating Its Hyperphosphorylation To Regulate Viral Propagation.J Virol. 2015 Jul;89(14):7385-400. doi: 10.1128/JVI.00567-15. Epub 2015 May 13. J Virol. 2015. PMID: 25972535 Free PMC article.
-
PACSIN2 Interacts with Nonstructural Protein 5A and Regulates Hepatitis C Virus Assembly.J Virol. 2020 Feb 14;94(5):e01531-19. doi: 10.1128/JVI.01531-19. Print 2020 Feb 14. J Virol. 2020. PMID: 31801866 Free PMC article.
-
Proprotein convertase subtilisin/kexin type 9 inhibits hepatitis C virus replication through interacting with NS5A.J Gen Virol. 2018 Jan;99(1):44-61. doi: 10.1099/jgv.0.000987. Epub 2017 Dec 13. J Gen Virol. 2018. PMID: 29235977
-
Phosphorylation of hepatitis C virus NS5A nonstructural protein: a new paradigm for phosphorylation-dependent viral RNA replication?Virology. 2007 Jul 20;364(1):1-9. doi: 10.1016/j.virol.2007.01.042. Epub 2007 Apr 2. Virology. 2007. PMID: 17400273 Review.
-
The non-structural 5A protein of hepatitis C virus.J Viral Hepat. 1999 Sep;6(5):343-56. doi: 10.1046/j.1365-2893.1999.00185.x. J Viral Hepat. 1999. PMID: 10607250 Review.
Cited by
-
TRIM25 Enhances the Antiviral Action of Zinc-Finger Antiviral Protein (ZAP).PLoS Pathog. 2017 Jan 6;13(1):e1006145. doi: 10.1371/journal.ppat.1006145. eCollection 2017 Jan. PLoS Pathog. 2017. PMID: 28060952 Free PMC article.
-
Hepatitis B Virus Stimulated Fibronectin Facilitates Viral Maintenance and Replication through Two Distinct Mechanisms.PLoS One. 2016 Mar 29;11(3):e0152721. doi: 10.1371/journal.pone.0152721. eCollection 2016. PLoS One. 2016. PMID: 27023403 Free PMC article.
-
Hepatitis C virus relies on lipoproteins for its life cycle.World J Gastroenterol. 2016 Feb 14;22(6):1953-65. doi: 10.3748/wjg.v22.i6.1953. World J Gastroenterol. 2016. PMID: 26877603 Free PMC article. Review.
-
Structural basis for autoinhibition and its relief of MOB1 in the Hippo pathway.Sci Rep. 2016 Jun 23;6:28488. doi: 10.1038/srep28488. Sci Rep. 2016. PMID: 27335147 Free PMC article.
-
The Spring α-Helix Coordinates Multiple Modes of HCV (Hepatitis C Virus) NS3 Helicase Action.J Biol Chem. 2016 Jul 8;291(28):14499-509. doi: 10.1074/jbc.M115.704379. Epub 2016 May 12. J Biol Chem. 2016. PMID: 27226535 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

