Breadth of neutralization and synergy of clinically relevant human monoclonal antibodies against HCV genotypes 1a, 1b, 2a, 2b, 2c, and 3a
- PMID: 25043937
- PMCID: PMC4415877
- DOI: 10.1002/hep.27298
Breadth of neutralization and synergy of clinically relevant human monoclonal antibodies against HCV genotypes 1a, 1b, 2a, 2b, 2c, and 3a
Abstract
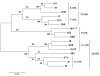
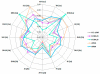
Human monoclonal antibodies (HMAbs) with neutralizing capabilities constitute potential immune-based treatments or prophylaxis against hepatitis C virus (HCV). However, lack of cell culture-derived HCV (HCVcc) harboring authentic envelope proteins (E1/E2) has hindered neutralization investigations across genotypes, subtypes, and isolates. We investigated the breadth of neutralization of 10 HMAbs with therapeutic potential against a panel of 16 JFH1-based HCVcc-expressing patient-derived Core-NS2 from genotypes 1a (strains H77, TN, and DH6), 1b (J4, DH1, and DH5), 2a (J6, JFH1, and T9), 2b (J8, DH8, and DH10), 2c (S83), and 3a (S52, DBN, and DH11). Virus stocks used for in vitro neutralization analysis contained authentic E1/E2, with the exception of full-length JFH1 that acquired the N417S substitution in E2. The 50% inhibition concentration (IC50) for each HMAb against the HCVcc panel was determined by dose-response neutralization assays in Huh7.5 cells with antibody concentrations ranging from 0.0012 to 100 μg/mL. Interestingly, IC50 values against the different HCVcc's exhibited large variations among the HMAbs, and only three HMAbs (HC-1AM, HC84.24, and AR4A) neutralized all 16 HCVcc recombinants. Furthermore, the IC50 values for a given HMAb varied greatly with the HCVcc strain, which supports the use of a diverse virus panel. In cooperation analyses, HMAbs HC84.24, AR3A, and, especially HC84.26, demonstrated synergistic effects towards the majority of the HCVcc's when combined individually with AR4A.
Conclusion: Through a neutralization analysis of 10 clinically relevant HMAbs against 16 JFH1-based Core-NS2 recombinants from genotypes 1a, 1b, 2a, 2b, 2c, and 3a, we identified at least three HMAbs with potent and broad neutralization potential. The neutralization synergism obtained when pooling the most potent HMAbs could have significant implications for developing novel strategies to treat and control HCV.
© 2014 by the American Association for the Study of Liver Diseases.
Figures


References
-
- Mohd HK, Groeger J, Flaxman AD, Wiersma ST. Global epidemiology of hepatitis C virus infection: new estimates of age-specific antibody to HCV seroprevalence. Hepatology. 2013;57(4):1333–1342. - PubMed
-
- Guillouche P, Feray C. Systematic review: anti-viral therapy of recurrent hepatitis C after liver transplantation. Aliment Pharmacol Ther. 2011;33(2):163–174. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
