The multiMiR R package and database: integration of microRNA-target interactions along with their disease and drug associations
- PMID: 25063298
- PMCID: PMC4176155
- DOI: 10.1093/nar/gku631
The multiMiR R package and database: integration of microRNA-target interactions along with their disease and drug associations
Abstract
microRNAs (miRNAs) regulate expression by promoting degradation or repressing translation of target transcripts. miRNA target sites have been catalogued in databases based on experimental validation and computational prediction using various algorithms. Several online resources provide collections of multiple databases but need to be imported into other software, such as R, for processing, tabulation, graphing and computation. Currently available miRNA target site packages in R are limited in the number of databases, types of databases and flexibility. We present multiMiR, a new miRNA-target interaction R package and database, which includes several novel features not available in existing R packages: (i) compilation of nearly 50 million records in human and mouse from 14 different databases, more than any other collection; (ii) expansion of databases to those based on disease annotation and drug microRNAresponse, in addition to many experimental and computational databases; and (iii) user-defined cutoffs for predicted binding strength to provide the most confident selection. Case studies are reported on various biomedical applications including mouse models of alcohol consumption, studies of chronic obstructive pulmonary disease in human subjects, and human cell line models of bladder cancer metastasis. We also demonstrate how multiMiR was used to generate testable hypotheses that were pursued experimentally.
© The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



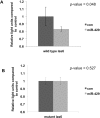
References
-
- Akbari Moqadam F., Pieters R., den Boer M.L. The hunting of targets: challenge in miRNA research. Leukemia. 2013;27:16–23. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

