metaseq: a Python package for integrative genome-wide analysis reveals relationships between chromatin insulators and associated nuclear mRNA
- PMID: 25063299
- PMCID: PMC4132753
- DOI: 10.1093/nar/gku644
metaseq: a Python package for integrative genome-wide analysis reveals relationships between chromatin insulators and associated nuclear mRNA
Abstract
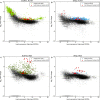
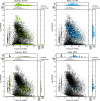
Here we introduce metaseq, a software library written in Python, which enables loading multiple genomic data formats into standard Python data structures and allows flexible, customized manipulation and visualization of data from high-throughput sequencing studies. We demonstrate its practical use by analyzing multiple datasets related to chromatin insulators, which are DNA-protein complexes proposed to organize the genome into distinct transcriptional domains. Recent studies in Drosophila and mammals have implicated RNA in the regulation of chromatin insulator activities. Moreover, the Drosophila RNA-binding protein Shep has been shown to antagonize gypsy insulator activity in a tissue-specific manner, but the precise role of RNA in this process remains unclear. Better understanding of chromatin insulator regulation requires integration of multiple datasets, including those from chromatin-binding, RNA-binding, and gene expression experiments. We use metaseq to integrate RIP- and ChIP-seq data for Shep and the core gypsy insulator protein Su(Hw) in two different cell types, along with publicly available ChIP-chip and RNA-seq data. Based on the metaseq-enabled analysis presented here, we propose a model where Shep associates with chromatin cotranscriptionally, then is recruited to insulator complexes in trans where it plays a negative role in insulator activity.
Published by Oxford University Press on behalf of Nucleic Acids Research 2014. This work is written by (a) US Government employee(s) and is in the public domain in the US.
Figures
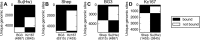



References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

