Toward a systems-level understanding of gene regulatory, protein interaction, and metabolic networks in cyanobacteria
- PMID: 25071821
- PMCID: PMC4079066
- DOI: 10.3389/fgene.2014.00191
Toward a systems-level understanding of gene regulatory, protein interaction, and metabolic networks in cyanobacteria
Abstract
Cyanobacteria are essential primary producers in marine ecosystems, playing an important role in both carbon and nitrogen cycles. In the last decade, various genome sequencing and metagenomic projects have generated large amounts of genetic data for cyanobacteria. This wealth of data provides researchers with a new basis for the study of molecular adaptation, ecology and evolution of cyanobacteria, as well as for developing biotechnological applications. It also facilitates the use of multiplex techniques, i.e., expression profiling by high-throughput technologies such as microarrays, RNA-seq, and proteomics. However, exploration and analysis of these data is challenging, and often requires advanced computational methods. Also, they need to be integrated into our existing framework of knowledge to use them to draw reliable biological conclusions. Here, systems biology provides important tools. Especially, the construction and analysis of molecular networks has emerged as a powerful systems-level framework, with which to integrate such data, and to better understand biological relevant processes in these organisms. In this review, we provide an overview of the advances and experimental approaches undertaken using multiplex data from genomic, transcriptomic, proteomic, and metabolomic studies in cyanobacteria. Furthermore, we summarize currently available web-based tools dedicated to cyanobacteria, i.e., CyanoBase, CyanoEXpress, ProPortal, Cyanorak, CyanoBIKE, and CINPER. Finally, we present a case study for the freshwater model cyanobacteria, Synechocystis sp. PCC6803, to show the power of meta-analysis, and the potential to extrapolate acquired knowledge to the ecologically important marine cyanobacteria genus, Prochlorococcus.
Keywords: cyanobacteria; meta-analysis; metabolic pathways; networks; systems biology.
Figures
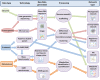


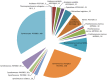
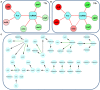
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

