An Illumina metabarcoding pipeline for fungi
- PMID: 25077016
- PMCID: PMC4113289
- DOI: 10.1002/ece3.1107
An Illumina metabarcoding pipeline for fungi
Abstract
High-throughput metabarcoding studies on fungi and other eukaryotic microorganisms are rapidly becoming more frequent and more complex, requiring researchers to handle ever increasing amounts of raw sequence data. Here, we provide a flexible pipeline for pruning and analyzing fungal barcode (ITS rDNA) data generated as paired-end reads on Illumina MiSeq sequencers. The pipeline presented includes specific steps fine-tuned for ITS, that are mostly missing from pipelines developed for prokaryotes. It (1) employs state of the art programs and follows best practices in fungal high-throughput metabarcoding; (2) consists of modules and scripts easily modifiable by the user to ensure maximum flexibility with regard to specific needs of a project or future methodological developments; and (3) is straightforward to use, also in classroom settings. We provide detailed descriptions and revision techniques for each step, thus giving the user maximum control over data treatment and avoiding a black-box approach. Employing this pipeline will improve and speed up the tedious and error-prone process of cleaning fungal Illumina metabarcoding data.
Keywords: community ecology; data pruning; high throughput; internal transcribed spacer rDNA; next-generation sequencing.
Figures
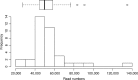
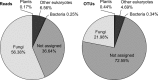
References
-
- Abarenkov K, Nilsson RH, Larsson K, Alexander IJ, Eberhardt U, Erland S, et al. The UNITE database for molecular identification of fungi – recent updates and future perspectives. New Phytol. 2010a;186:281–285. - PubMed
-
- Abarenkov K, Tedersoo L, Nilsson RH, Vellak K, Saar I, Veldre V, et al. PlutoF – a Web Based Workbench for ecological and taxonomic research, with an online implementation for fungal ITS sequences. Evol. Bioinform. Online. 2010b;6:189–196.
-
- Bengtsson-Palme J, Ryberg M, Hartmann M, Branco S, Wang Z, Godhe A, et al. Improved software detection and extraction of ITS1 and ITS2 from ribosomal ITS sequences of fungi and other eukaryotes for analysis of environmental sequencing data. Methods Ecol. Evol. 2013;4:914–919.
LinkOut - more resources
Full Text Sources
Other Literature Sources

