Oligomeric state of purified transient receptor potential melastatin-1 (TRPM1), a protein essential for dim light vision
- PMID: 25112866
- PMCID: PMC4175340
- DOI: 10.1074/jbc.M114.593780
Oligomeric state of purified transient receptor potential melastatin-1 (TRPM1), a protein essential for dim light vision
Abstract
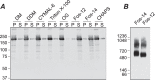
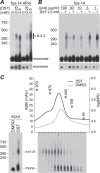
Transient receptor potential melastatin-1 (TRPM1) is essential for the light-induced depolarization of retinal ON bipolar cells. TRPM1 likely forms a multimeric channel complex, although almost nothing is known about the structure or subunit composition of channels formed by TRPM1 or any of its close relatives. Recombinant TRPM1 was robustly expressed in insect cells, but only a small fraction was localized to the plasma membrane. Similar intracellular localization was observed when TRPM1 was heterologously expressed in mammalian cells. TRPM1 was affinity-purified from Sf9 cells and complexed with amphipol, followed by detergent removal. In blue native gels and size exclusion chromatography, TRPM1 migrated with a mobility consistent with detergent- or amphipol-bound dimers. Cross-linking experiments were also consistent with a dimeric subunit stoichiometry, and cryoelectron microscopy and single particle analysis without symmetry imposition yielded a model with approximate 2-fold symmetrical features. Finally, electron microscopy of TRPM1-antibody complexes revealed a large particle that can accommodate TRPM1 and two antibody molecules. Taken together, these data indicate that purified TRPM1 is mostly dimeric. The three-dimensional structure of TRPM1 dimers is characterized by a small putative transmembrane domain and a larger domain with a hollow cavity. Blue native gels of solubilized mouse retina indicate that TRPM1 is present in two distinct complexes: one similar in size to the recombinant protein and one much larger. Because dimers are likely not functional ion channels, these results suggest that additional partner subunits participate in forming the transduction channel required for dim light vision and the ON pathway.
Keywords: Cryoelectron Microscopy; G Protein Signaling; Ion Channel; Membrane Protein; Transient Receptor Potential Channels (TRP Channels); Vision.
© 2014 by The American Society for Biochemistry and Molecular Biology, Inc.
Figures








Similar articles
-
A Large Endoplasmic Reticulum-Resident Pool of TRPM1 in Retinal ON-Bipolar Cells.eNeuro. 2018 Jul 4;5(3):ENEURO.0143-18.2018. doi: 10.1523/ENEURO.0143-18.2018. eCollection 2018 May-Jun. eNeuro. 2018. PMID: 30027108 Free PMC article.
-
Characterization of the part of N-terminal PIP2 binding site of the TRPM1 channel.Biophys Chem. 2015 Dec;207:135-42. doi: 10.1016/j.bpc.2015.10.005. Epub 2015 Oct 27. Biophys Chem. 2015. PMID: 26544986
-
G-protein-mediated inhibition of the Trp channel TRPM1 requires the Gβγ dimer.Proc Natl Acad Sci U S A. 2012 May 29;109(22):8752-7. doi: 10.1073/pnas.1117433109. Epub 2012 May 14. Proc Natl Acad Sci U S A. 2012. PMID: 22586107 Free PMC article.
-
TRPM1: a vertebrate TRP channel responsible for retinal ON bipolar function.Cell Calcium. 2010 Aug-Sep;48(2-3):95-101. doi: 10.1016/j.ceca.2010.08.004. Epub 2010 Sep 16. Cell Calcium. 2010. PMID: 20846719 Review.
-
TRPM1.Handb Exp Pharmacol. 2014;222:387-402. doi: 10.1007/978-3-642-54215-2_15. Handb Exp Pharmacol. 2014. PMID: 24756714 Review.
Cited by
-
Therapeutic Antibodies Targeting Potassium Ion Channels.Handb Exp Pharmacol. 2021;267:507-545. doi: 10.1007/164_2021_464. Handb Exp Pharmacol. 2021. PMID: 33963460
-
A Large Endoplasmic Reticulum-Resident Pool of TRPM1 in Retinal ON-Bipolar Cells.eNeuro. 2018 Jul 4;5(3):ENEURO.0143-18.2018. doi: 10.1523/ENEURO.0143-18.2018. eCollection 2018 May-Jun. eNeuro. 2018. PMID: 30027108 Free PMC article.
-
Critical Role for Phosphatidylinositol-3 Kinase Vps34/PIK3C3 in ON-Bipolar Cells.Invest Ophthalmol Vis Sci. 2019 Jul 1;60(8):2861-2874. doi: 10.1167/iovs.19-26586. Invest Ophthalmol Vis Sci. 2019. PMID: 31260037 Free PMC article.
-
Long-term follow-up of retinal function and structure in TRPM1-associated complete congenital stationary night blindness.Mol Vis. 2019 Dec 19;25:851-858. eCollection 2019. Mol Vis. 2019. PMID: 31908403 Free PMC article.
-
The Transduction Cascade in Retinal ON-Bipolar Cells: Signal Processing and Disease.Annu Rev Vis Sci. 2017 Sep 15;3:25-51. doi: 10.1146/annurev-vision-102016-061338. Epub 2017 Jul 17. Annu Rev Vis Sci. 2017. PMID: 28715957 Free PMC article. Review.
References
-
- Lukasiewicz P. D. (2005) Synaptic mechanisms that shape visual signaling at the inner retina. Prog. Brain Res. 147, 205–218 - PubMed
-
- Masu M., Iwakabe H., Tagawa Y., Miyoshi T., Yamashita M., Fukuda Y., Sasaki H., Hiroi K., Nakamura Y., Shigemoto R., Takada M., Nakamura K., Nakao K., Katsuki M., Nakanishi S. (1995) Specific deficit of the ON response in visual transmission by targeted disruption of the mGluR6 gene. Cell 80, 757–765 - PubMed
-
- Nakajima Y., Iwakabe H., Akazawa C., Nawa H., Shigemoto R., Mizuno N., Nakanishi S. (1993) Molecular characterization of a novel retinal metabotropic glutamate receptor mGluR6 with a high agonist selectivity for l-2-amino-4-phosphonobutyrate. J. Biol. Chem. 268, 11868–11873 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
- R01 EY07981/EY/NEI NIH HHS/United States
- R01 EY11900/EY/NEI NIH HHS/United States
- K12 GM084897/GM/NIGMS NIH HHS/United States
- R01 GM066099/GM/NIGMS NIH HHS/United States
- R01 EY011900/EY/NEI NIH HHS/United States
- F32 EY200672/EY/NEI NIH HHS/United States
- T32 DK007696/DK/NIDDK NIH HHS/United States
- P30 CA125123/CA/NCI NIH HHS/United States
- P30 EY002520/EY/NEI NIH HHS/United States
- F32 EY024815/EY/NEI NIH HHS/United States
- T32 EY007102/EY/NEI NIH HHS/United States
- F32 EY020067/EY/NEI NIH HHS/United States
- R01 EY007981/EY/NEI NIH HHS/United States
- P41 GM103832/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

