Characterizing the proton loading site in cytochrome c oxidase
- PMID: 25114210
- PMCID: PMC4151712
- DOI: 10.1073/pnas.1407187111
Characterizing the proton loading site in cytochrome c oxidase
Abstract
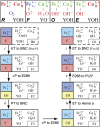
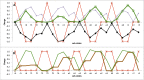
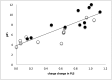
Cytochrome c oxidase (CcO) uses the energy released by reduction of O2 to H2O to drive eight charges from the high pH to low pH side of the membrane, increasing the electrochemical gradient. Four electrons and protons are used for chemistry, while four more protons are pumped. Proton pumping requires that residues on a pathway change proton affinity through the reaction cycle to load and then release protons. The protonation states of all residues in CcO are determined in MultiConformational Continuum Electrostatics simulations with the protonation and redox states of heme a, a3, Cu(B), Y288, and E286 used to define the catalytic cycle. One proton is found to be loaded and released from residues identified as the proton loading site (PLS) on the P-side of the protein in each of the four CcO redox states. Thus, the same proton pumping mechanism can be used each time CcO is reduced. Calculations with structures of Rhodobacter sphaeroides, Paracoccus denitrificans, and bovine CcO derived by crystallography and molecular dynamics show the PLS functions similarly in different CcO species. The PLS is a cluster rather than a single residue, as different structures show 1-4 residues load and release protons. However, the proton affinity of the heme a3 propionic acids primarily determines the number of protons loaded into the PLS; if their proton affinity is too low, less than one proton is loaded.
Keywords: MCCE; bioenergetics; pKa; proton transfer.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Identifying the proton loading site cluster in the ba3 cytochrome c oxidase that loads and traps protons.Biochim Biophys Acta Bioenerg. 2020 Oct 1;1861(10):148239. doi: 10.1016/j.bbabio.2020.148239. Epub 2020 Jun 10. Biochim Biophys Acta Bioenerg. 2020. PMID: 32531221
-
Calculated proton uptake on anaerobic reduction of cytochrome C oxidase: is the reaction electroneutral?Biochemistry. 2006 Jul 4;45(26):7959-75. doi: 10.1021/bi052183d. Biochemistry. 2006. PMID: 16800622 Free PMC article.
-
Similarity of cytochrome c oxidases in different organisms.Proteins. 2010 Sep;78(12):2691-8. doi: 10.1002/prot.22783. Proteins. 2010. PMID: 20589635 Free PMC article.
-
Respiratory conservation of energy with dioxygen: cytochrome C oxidase.Met Ions Life Sci. 2015;15:89-130. doi: 10.1007/978-3-319-12415-5_4. Met Ions Life Sci. 2015. PMID: 25707467 Review.
-
Variable proton-pumping stoichiometry in structural variants of cytochrome c oxidase.Biochim Biophys Acta. 2010 Jun-Jul;1797(6-7):710-23. doi: 10.1016/j.bbabio.2010.02.020. Epub 2010 Feb 23. Biochim Biophys Acta. 2010. PMID: 20184858 Review.
Cited by
-
Multiscale simulations reveal key features of the proton-pumping mechanism in cytochrome c oxidase.Proc Natl Acad Sci U S A. 2016 Jul 5;113(27):7420-5. doi: 10.1073/pnas.1601982113. Epub 2016 Jun 23. Proc Natl Acad Sci U S A. 2016. PMID: 27339133 Free PMC article.
-
Redox-Driven Proton Pumps of the Respiratory Chain.Biophys J. 2018 Sep 4;115(5):830-840. doi: 10.1016/j.bpj.2018.07.022. Epub 2018 Aug 2. Biophys J. 2018. PMID: 30119834 Free PMC article.
-
Long-range charge transfer mechanism of the III2IV2 mycobacterial supercomplex.Nat Commun. 2024 Jun 20;15(1):5276. doi: 10.1038/s41467-024-49628-9. Nat Commun. 2024. PMID: 38902248 Free PMC article.
-
The electron-proton bottleneck of photosynthetic oxygen evolution.Nature. 2023 May;617(7961):623-628. doi: 10.1038/s41586-023-06008-5. Epub 2023 May 3. Nature. 2023. PMID: 37138082 Free PMC article.
-
Mutation of a single residue in the ba3 oxidase specifically impairs protonation of the pump site.Proc Natl Acad Sci U S A. 2015 Mar 17;112(11):3397-402. doi: 10.1073/pnas.1422434112. Epub 2015 Mar 2. Proc Natl Acad Sci U S A. 2015. PMID: 25733886 Free PMC article.
References
-
- Mitchell P, Moyle J. Stoichiometry of proton translocation through the respiratory chain and adenosine triphosphatase systems of rat liver mitochondria. Nature. 1965;208(5006):147–151. - PubMed
-
- Babcock GT, Wikström M. Oxygen activation and the conservation of energy in cell respiration. Nature. 1992;356(6367):301–309. - PubMed
-
- Kaila VR, Verkhovsky MI, Wikström M. Proton-coupled electron transfer in cytochrome oxidase. Chem Rev. 2010;110(12):7062–7081. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

