Frequent changes in expression profile and accelerated sequence evolution of duplicated imprinted genes in arabidopsis
- PMID: 25115008
- PMCID: PMC4122942
- DOI: 10.1093/gbe/evu144
Frequent changes in expression profile and accelerated sequence evolution of duplicated imprinted genes in arabidopsis
Abstract
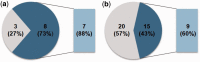
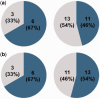
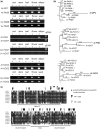
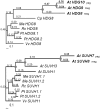
Eukaryotic genomes have large numbers of duplicated genes that can evolve new functions or expression patterns by changes in coding and regulatory sequences, referred to as neofunctionalization. In flowering plants, some duplicated genes are imprinted in the endosperm, where only one allele is expressed depending on its parental origin. We found that 125 imprinted genes in Arabidopsis arose from gene duplication events during the evolution of the Brassicales. Analyses of 46 gene pairs duplicated by an ancient whole-genome duplication (alpha WGD) indicated that many imprinted genes show an accelerated rate of amino acid changes compared with their paralogs. Analyses of microarray expression data from 63 organ types and developmental stages indicated that many imprinted genes have expression patterns restricted to flowers and/or seeds in contrast to their broadly expressed paralogs. Assays of expression in orthologs from outgroup species revealed that some imprinted genes have acquired an organ-specific expression pattern restricted to flowers and/or seeds. The changes in expression pattern and the accelerated sequence evolution in the imprinted genes suggest that some of them may have undergone neofunctionalization. The imprinted genes MPC, HOMEODOMAIN GLABROUS6 (HDG6), and HDG3 are particularly interesting cases that have different functions from their paralogs. This study indicates that a large number of imprinted genes in Arabidopsis are evolutionarily recent duplicates and that many of them show changes in expression profiles and accelerated sequence evolution. Acquisition of imprinting is a mode of duplicate gene divergence in plants that is more common than previously thought.
Figures

Similar articles
-
Extensive divergence in alternative splicing patterns after gene and genome duplication during the evolutionary history of Arabidopsis.Mol Biol Evol. 2010 Jul;27(7):1686-97. doi: 10.1093/molbev/msq054. Epub 2010 Feb 25. Mol Biol Evol. 2010. PMID: 20185454
-
Protein subcellular relocalization of duplicated genes in Arabidopsis.Genome Biol Evol. 2014 Sep 4;6(9):2501-15. doi: 10.1093/gbe/evu191. Genome Biol Evol. 2014. PMID: 25193306 Free PMC article.
-
Dramatic change in function and expression pattern of a gene duplicated by polyploidy created a paternal effect gene in the Brassicaceae.Mol Biol Evol. 2010 Dec;27(12):2817-28. doi: 10.1093/molbev/msq169. Epub 2010 Jul 8. Mol Biol Evol. 2010. PMID: 20616146
-
Gene duplication within the Green Lineage: the case of TEL genes.J Exp Bot. 2012 Sep;63(14):5061-77. doi: 10.1093/jxb/ers181. Epub 2012 Aug 3. J Exp Bot. 2012. PMID: 22865910 Review.
-
Epigenetic mechanisms underlying genomic imprinting in plants.Annu Rev Plant Biol. 2012;63:331-52. doi: 10.1146/annurev-arplant-042811-105514. Epub 2012 Jan 30. Annu Rev Plant Biol. 2012. PMID: 22404470 Review.
Cited by
-
Large-Scale Evolutionary Analysis of Genes and Supergene Clusters from Terpenoid Modular Pathways Provides Insights into Metabolic Diversification in Flowering Plants.PLoS One. 2015 Jun 5;10(6):e0128808. doi: 10.1371/journal.pone.0128808. eCollection 2015. PLoS One. 2015. PMID: 26046541 Free PMC article.
-
Concerted Divergence after Gene Duplication in Polycomb Repressive Complexes.Plant Physiol. 2017 Jun;174(2):1192-1204. doi: 10.1104/pp.16.01983. Epub 2017 Apr 28. Plant Physiol. 2017. PMID: 28455403 Free PMC article.
-
Whole-genome duplication and molecular evolution in Cornus L. (Cornaceae) - Insights from transcriptome sequences.PLoS One. 2017 Feb 22;12(2):e0171361. doi: 10.1371/journal.pone.0171361. eCollection 2017. PLoS One. 2017. PMID: 28225773 Free PMC article.
-
Paternally Expressed Imprinted Genes under Positive Darwinian Selection in Arabidopsis thaliana.Mol Biol Evol. 2019 Jun 1;36(6):1239-1253. doi: 10.1093/molbev/msz063. Mol Biol Evol. 2019. PMID: 30913563 Free PMC article.
-
Genomic Imprinting Was Evolutionarily Conserved during Wheat Polyploidization.Plant Cell. 2018 Jan;30(1):37-47. doi: 10.1105/tpc.17.00837. Epub 2018 Jan 3. Plant Cell. 2018. PMID: 29298834 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

