IglC and PdpA are important for promoting Francisella invasion and intracellular growth in epithelial cells
- PMID: 25115488
- PMCID: PMC4130613
- DOI: 10.1371/journal.pone.0104881
IglC and PdpA are important for promoting Francisella invasion and intracellular growth in epithelial cells
Abstract
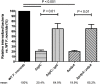
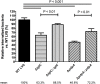
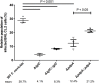
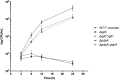
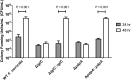
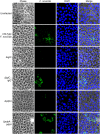
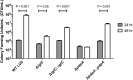
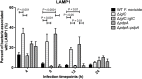
The highly infectious bacteria, Francisella tularensis, colonize a variety of organs and replicate within both phagocytic as well as non-phagocytic cells, to cause the disease tularemia. These microbes contain a conserved cluster of important virulence genes referred to as the Francisella Pathogenicity Island (FPI). Two of the most characterized FPI genes, iglC and pdpA, play a central role in bacterial survival and proliferation within phagocytes, but do not influence bacterial internalization. Yet, their involvement in non-phagocytic epithelial cell infections remains unexplored. To examine the functions of IglC and PdpA on bacterial invasion and replication during epithelial cell infections, we infected liver and lung epithelial cells with F. novicida and F. tularensis 'Type B' Live Vaccine Strain (LVS) deletion mutants (ΔiglC and ΔpdpA) as well as their respective gene complements. We found that deletion of either gene significantly reduced their ability to invade and replicate in epithelial cells. Gene complementation of iglC and pdpA partially rescued bacterial invasion and intracellular growth. Additionally, substantial LAMP1-association with both deletion mutants was observed up to 12 h suggesting that the absence of IglC and PdpA caused deficiencies in their ability to dissociate from LAMP1-positive Francisella Containing Vacuoles (FCVs). This work provides the first evidence that IglC and PdpA are important pathogenic factors for invasion and intracellular growth of Francisella in epithelial cells, and further highlights the discrete mechanisms involved in Francisella infections between phagocytic and non-phagocytic cells.
Conflict of interest statement
Figures
Similar articles
-
The Francisella tularensis pathogenicity island protein IglC and its regulator MglA are essential for modulating phagosome biogenesis and subsequent bacterial escape into the cytoplasm.Cell Microbiol. 2005 Jul;7(7):969-79. doi: 10.1111/j.1462-5822.2005.00526.x. Cell Microbiol. 2005. PMID: 15953029
-
Characterization of the pathogenicity island protein PdpA and its role in the virulence of Francisella novicida.Microbiology (Reading). 2009 May;155(Pt 5):1489-1497. doi: 10.1099/mic.0.025379-0. Epub 2009 Apr 16. Microbiology (Reading). 2009. PMID: 19372153 Free PMC article.
-
A Francisella novicida pdpA mutant exhibits limited intracellular replication and remains associated with the lysosomal marker LAMP-1.Microbiology (Reading). 2009 May;155(Pt 5):1498-1504. doi: 10.1099/mic.0.025445-0. Epub 2009 Apr 16. Microbiology (Reading). 2009. PMID: 19372155 Free PMC article.
-
Molecular and genetic basis of pathogenesis in Francisella tularensis.Ann N Y Acad Sci. 2007 Jun;1105:138-59. doi: 10.1196/annals.1409.010. Epub 2007 Mar 29. Ann N Y Acad Sci. 2007. PMID: 17395737 Review.
-
Loops and networks in control of Francisella tularensis virulence.Future Microbiol. 2009 Aug;4(6):713-29. doi: 10.2217/fmb.09.37. Future Microbiol. 2009. PMID: 19659427 Review.
Cited by
-
Entry of Francisella tularensis into Murine B Cells: The Role of B Cell Receptors and Complement Receptors.PLoS One. 2015 Jul 10;10(7):e0132571. doi: 10.1371/journal.pone.0132571. eCollection 2015. PLoS One. 2015. PMID: 26161475 Free PMC article.
-
Fosmidomycin, an inhibitor of isoprenoid synthesis, induces persistence in Chlamydia by inhibiting peptidoglycan assembly.PLoS Pathog. 2019 Oct 17;15(10):e1008078. doi: 10.1371/journal.ppat.1008078. eCollection 2019 Oct. PLoS Pathog. 2019. PMID: 31622442 Free PMC article.
-
Transcription Elongation Factor GreA Plays a Key Role in Cellular Invasion and Virulence of Francisella tularensis subsp. novicida.Sci Rep. 2018 May 2;8(1):6895. doi: 10.1038/s41598-018-25271-5. Sci Rep. 2018. PMID: 29720697 Free PMC article.
-
Dual Role of Mechanisms Involved in Resistance to Predation by Protozoa and Virulence to Humans.Front Microbiol. 2018 May 17;9:1017. doi: 10.3389/fmicb.2018.01017. eCollection 2018. Front Microbiol. 2018. PMID: 29867902 Free PMC article. Review.
-
Microinjection of Francisella tularensis and Listeria monocytogenes reveals the importance of bacterial and host factors for successful replication.Infect Immun. 2015 Aug;83(8):3233-42. doi: 10.1128/IAI.00416-15. Epub 2015 Jun 1. Infect Immun. 2015. PMID: 26034213 Free PMC article.
References
-
- Sjöstedt AB (2005) Francisella. In The Proteobacteria, Part B. In: Brenner DJ, Staley JT, Garrity GM, editors. Bergey’s Manual of Systematic Bacteriology. 2 ed. New York, NY: Springer. 200–210.
-
- Santic M, Al-Khodor S, Abu Kwaik Y (2010) Cell biology and molecular ecology of Francisella tularensis . Cell Microbiol 12: 129–139. - PubMed
-
- Huber B, Escudero R, Busse HJ, Seibold E, Scholz HC, et al. (2010) Description of Francisella hispaniensis sp. nov., isolated from human blood, reclassification of Francisella novicida (Larson, et al. 1955) Olsufiev, et al. 1959 as Francisella tularensis subsp. novicida comb. nov. and emended description of the genus Francisella. Int J Syst Evol Microbiol 60: 1887–1896. - PubMed
-
- Busse HJ, Huber B, Anda P, Escudero R, Scholz HC, et al. (2010) Objections to the transfer of Francisella novicida to the subspecies rank of Francisella tularensis - response to Johansson, et al. Int J Syst Evol Microbiol 60: 1718–1720. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

