Complex communities of small protists and unexpected occurrence of typical marine lineages in shallow freshwater systems
- PMID: 25115943
- PMCID: PMC4232972
- DOI: 10.1111/1462-2920.12591
Complex communities of small protists and unexpected occurrence of typical marine lineages in shallow freshwater systems
Abstract
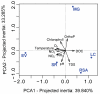
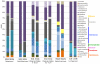
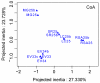
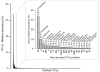
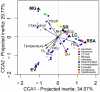
Although inland water bodies are more heterogeneous and sensitive to environmental variation than oceans, the diversity of small protists in these ecosystems is much less well known. Some molecular surveys of lakes exist, but little information is available from smaller, shallower and often ephemeral freshwater systems, despite their global distribution and ecological importance. We carried out a comparative study based on massive pyrosequencing of amplified 18S rRNA gene fragments of protists in the 0.2-5 μm size range in one brook and four shallow ponds located in the Natural Regional Park of the Chevreuse Valley, France. Our study revealed a wide diversity of small protists, with 812 stringently defined operational taxonomic units (OTUs) belonging to the recognized eukaryotic supergroups (SAR--Stramenopiles, Alveolata, Rhizaria--Archaeplastida, Excavata, Amoebozoa, Opisthokonta) and to groups of unresolved phylogenetic position (Cryptophyta, Haptophyta, Centrohelida, Katablepharida, Telonemida, Apusozoa). Some OTUs represented deep-branching lineages (Cryptomycota, Aphelida, Colpodellida, Tremulida, clade-10 Cercozoa, HAP-1 Haptophyta). We identified several lineages previously thought to be marine including, in addition to MAST-2 and MAST-12, already detected in freshwater, MAST-3 and possibly MAST-6. Protist community structures were different in the five ecosystems. These differences did not correlate with geographical distances, but seemed to be influenced by environmental parameters.
© 2014 Society for Applied Microbiology and John Wiley & Sons Ltd.
Figures
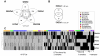

References
-
- Baas-Becking L. Geobiologie of inleiding tot de milieukunde. W.P. Van Stock; Zoon, Hague, Netherlands: 1934.
-
- Bass D, Chao EE-Y, Nikolaev S, Yabuki A, Ishida K, Berney C, et al. Phylogeny of Novel Naked Filose and Reticulose Cercozoa: Granofilosea cl. n. and Proteomyxidea Revised. Protist. 2009;160:75–109. - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

