A long noncoding RNA protects the heart from pathological hypertrophy
- PMID: 25119045
- PMCID: PMC4184960
- DOI: 10.1038/nature13596
A long noncoding RNA protects the heart from pathological hypertrophy
Abstract
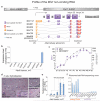
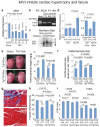
The role of long noncoding RNA (lncRNA) in adult hearts is unknown; also unclear is how lncRNA modulates nucleosome remodelling. An estimated 70% of mouse genes undergo antisense transcription, including myosin heavy chain 7 (Myh7), which encodes molecular motor proteins for heart contraction. Here we identify a cluster of lncRNA transcripts from Myh7 loci and demonstrate a new lncRNA-chromatin mechanism for heart failure. In mice, these transcripts, which we named myosin heavy-chain-associated RNA transcripts (Myheart, or Mhrt), are cardiac-specific and abundant in adult hearts. Pathological stress activates the Brg1-Hdac-Parp chromatin repressor complex to inhibit Mhrt transcription in the heart. Such stress-induced Mhrt repression is essential for cardiomyopathy to develop: restoring Mhrt to the pre-stress level protects the heart from hypertrophy and failure. Mhrt antagonizes the function of Brg1, a chromatin-remodelling factor that is activated by stress to trigger aberrant gene expression and cardiac myopathy. Mhrt prevents Brg1 from recognizing its genomic DNA targets, thus inhibiting chromatin targeting and gene regulation by Brg1. It does so by binding to the helicase domain of Brg1, a domain that is crucial for tethering Brg1 to chromatinized DNA targets. Brg1 helicase has dual nucleic-acid-binding specificities: it is capable of binding lncRNA (Mhrt) and chromatinized--but not naked--DNA. This dual-binding feature of helicase enables a competitive inhibition mechanism by which Mhrt sequesters Brg1 from its genomic DNA targets to prevent chromatin remodelling. A Mhrt-Brg1 feedback circuit is thus crucial for heart function. Human MHRT also originates from MYH7 loci and is repressed in various types of myopathic hearts, suggesting a conserved lncRNA mechanism in human cardiomyopathy. Our studies identify a cardioprotective lncRNA, define a new targeting mechanism for ATP-dependent chromatin-remodelling factors, and establish a new paradigm for lncRNA-chromatin interaction.
Figures



Comment in
-
An epigenetic "LINK(RNA)" to pathological cardiac hypertrophy.Cell Metab. 2014 Oct 7;20(4):555-7. doi: 10.1016/j.cmet.2014.09.011. Cell Metab. 2014. PMID: 25295782 Free PMC article.
-
Dawn of the Epi-LncRNAs: new path from Myheart.Circ Res. 2015 Jan 16;116(2):235-6. doi: 10.1161/CIRCRESAHA.114.305490. Circ Res. 2015. PMID: 25593274 Free PMC article.
-
Myheart hits the core of chromatin.Cell Cycle. 2015;14(6):787-8. doi: 10.1080/15384101.2015.1010963. Cell Cycle. 2015. PMID: 25668574 Free PMC article. No abstract available.
Similar articles
-
Chromatin regulation by Brg1 underlies heart muscle development and disease.Nature. 2010 Jul 1;466(7302):62-7. doi: 10.1038/nature09130. Nature. 2010. PMID: 20596014 Free PMC article.
-
Epigenetic response to environmental stress: Assembly of BRG1-G9a/GLP-DNMT3 repressive chromatin complex on Myh6 promoter in pathologically stressed hearts.Biochim Biophys Acta. 2016 Jul;1863(7 Pt B):1772-81. doi: 10.1016/j.bbamcr.2016.03.002. Epub 2016 Mar 4. Biochim Biophys Acta. 2016. PMID: 26952936 Free PMC article.
-
T3 Critically Affects the Mhrt/Brg1 Axis to Regulate the Cardiac MHC Switch: Role of an Epigenetic Cross-Talk.Cells. 2020 Sep 24;9(10):2155. doi: 10.3390/cells9102155. Cells. 2020. PMID: 32987653 Free PMC article.
-
Epigenetic and lncRNA regulation of cardiac pathophysiology.Biochim Biophys Acta. 2016 Jul;1863(7 Pt B):1767-71. doi: 10.1016/j.bbamcr.2016.03.005. Epub 2016 Mar 9. Biochim Biophys Acta. 2016. PMID: 26969820 Free PMC article. Review.
-
Chromatin modifications remodel cardiac gene expression.Cardiovasc Res. 2014 Jul 1;103(1):7-16. doi: 10.1093/cvr/cvu122. Epub 2014 May 8. Cardiovasc Res. 2014. PMID: 24812277 Review.
Cited by
-
A myocardin-adjacent lncRNA balances SRF-dependent gene transcription in the heart.Genes Dev. 2021 Jun;35(11-12):835-840. doi: 10.1101/gad.348304.121. Epub 2021 May 13. Genes Dev. 2021. PMID: 33985971 Free PMC article.
-
(MYO)SLIDing Our Way Into the Vascular Pool of Long Noncoding RNAs.Arterioscler Thromb Vasc Biol. 2016 Oct;36(10):2033-4. doi: 10.1161/ATVBAHA.116.308173. Arterioscler Thromb Vasc Biol. 2016. PMID: 27655778 Free PMC article.
-
Heart failure: advanced development in genetics and epigenetics.Biomed Res Int. 2015;2015:352734. doi: 10.1155/2015/352734. Epub 2015 Apr 9. Biomed Res Int. 2015. PMID: 25949994 Free PMC article. Review.
-
Readers, writers, and erasers: chromatin as the whiteboard of heart disease.Circ Res. 2015 Mar 27;116(7):1245-53. doi: 10.1161/CIRCRESAHA.116.303630. Circ Res. 2015. PMID: 25814685 Free PMC article. Review.
-
Deletion of the muscle enriched lncRNA Oip5os1 induces atrial dysfunction in male mice with diabetes.Physiol Rep. 2023 Dec;11(23):e15869. doi: 10.14814/phy2.15869. Physiol Rep. 2023. PMID: 38054572 Free PMC article.
References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
- HL109512/HL/NHLBI NIH HHS/United States
- HL71140/HL/NHLBI NIH HHS/United States
- R01 HL105194/HL/NHLBI NIH HHS/United States
- R01 HL111770/HL/NHLBI NIH HHS/United States
- HL116997/HL/NHLBI NIH HHS/United States
- HL121197/HL/NHLBI NIH HHS/United States
- HL118087/HL/NHLBI NIH HHS/United States
- HL78931/HL/NHLBI NIH HHS/United States
- R01 HL118087/HL/NHLBI NIH HHS/United States
- R01 HL116997/HL/NHLBI NIH HHS/United States
- HL105194/HL/NHLBI NIH HHS/United States
- R01 HL121197/HL/NHLBI NIH HHS/United States
- HL111770/HL/NHLBI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

